Sulfolobus Spindle-Shaped Virus 1 Contains Glycosylated Capsid Proteins, a Cellular Chromatin Protein, and Host-Derived Lipids
- PMID: 26355093
- PMCID: PMC4645638
- DOI: 10.1128/JVI.02270-15
Sulfolobus Spindle-Shaped Virus 1 Contains Glycosylated Capsid Proteins, a Cellular Chromatin Protein, and Host-Derived Lipids
Abstract
Geothermal and hypersaline environments are rich in virus-like particles, among which spindle-shaped morphotypes dominate. Currently, viruses with spindle- or lemon-shaped virions are exclusive to Archaea and belong to two distinct viral families. The larger of the two families, the Fuselloviridae, comprises tail-less, spindle-shaped viruses, which infect hosts from phylogenetically distant archaeal lineages. Sulfolobus spindle-shaped virus 1 (SSV1) is the best known member of the family and was one of the first hyperthermophilic archaeal viruses to be isolated. SSV1 is an attractive model for understanding virus-host interactions in Archaea; however, the constituents and architecture of SSV1 particles remain only partially characterized. Here, we have conducted an extensive biochemical characterization of highly purified SSV1 virions and identified four virus-encoded structural proteins, VP1 to VP4, as well as one DNA-binding protein of cellular origin. The virion proteins VP1, VP3, and VP4 undergo posttranslational modification by glycosylation, seemingly at multiple sites. VP1 is also proteolytically processed. In addition to the viral DNA-binding protein VP2, we show that viral particles contain the Sulfolobus solfataricus chromatin protein Sso7d. Finally, we provide evidence indicating that SSV1 virions contain glycerol dibiphytanyl glycerol tetraether (GDGT) lipids, resolving a long-standing debate on the presence of lipids within SSV1 virions. A comparison of the contents of lipids isolated from the virus and its host cell suggests that GDGTs are acquired by the virus in a selective manner from the host cytoplasmic membrane, likely during progeny egress.
Importance: Although spindle-shaped viruses represent one of the most prominent viral groups in Archaea, structural data on their virion constituents and architecture still are scarce. The comprehensive biochemical characterization of the hyperthermophilic virus SSV1 presented here brings novel and significant insights into the organization and architecture of spindle-shaped virions. The obtained data permit the comparison between spindle-shaped viruses residing in widely different ecological niches, improving our understanding of the adaptation of viruses with unusual morphotypes to extreme environmental conditions.
Copyright © 2015, American Society for Microbiology. All Rights Reserved.
Figures
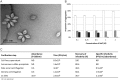



References
-
- Atanasova NS, Senčilo A, Pietilä MK, Roine E, Oksanen HM, Bamford DH. 2015. Comparison of lipid-containing bacterial and archaeal viruses. Adv Virus Res 92:1–61. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

