Relationships of wild and domesticated rices (Oryza AA genome species) based upon whole chloroplast genome sequences
- PMID: 26355750
- PMCID: PMC4564799
- DOI: 10.1038/srep13957
Relationships of wild and domesticated rices (Oryza AA genome species) based upon whole chloroplast genome sequences
Abstract
Rice is the most important crop in the world, acting as the staple food for over half of the world's population. The evolutionary relationships of cultivated rice and its wild relatives have remained contentious and inconclusive. Here we report on the use of whole chloroplast sequences to elucidate the evolutionary and phylogenetic relationships in the AA genome Oryza species, representing the primary gene pool of rice. This is the first study that has produced a well resolved and strongly supported phylogeny of the AA genome species. The pan tropical distribution of these rice relatives was found to be explained by long distance dispersal within the last million years. The analysis resulted in a clustering pattern that showed strong geographical differentiation. The species were defined in two primary clades with a South American/African clade with two species, O glumaepatula and O longistaminata, distinguished from all other species. The largest clade was comprised of an Australian clade including newly identified taxa and the African and Asian clades. This refined knowledge of the relationships between cultivated rice and the related wild species provides a strong foundation for more targeted use of wild genetic resources in rice improvement and efforts to ensure their conservation.
Figures

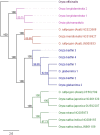
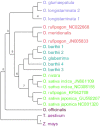
References
-
- Chang T.-T. The origin, evolution, cultivation, dissemination, and diversification of Asian and African rices. Euphytica 25, 425–441 (1976).
-
- Zhu T. et al. Phylogenetic relationships and genome divergence among the AA- genome species of the genus Oryza as revealed by 53 nuclear genes and 16 intergenic regions. Mol. Phylogenet. Evol. 70, 348–361 (2014). - PubMed
-
- Zhu Q. & Ge S. Phylogenetic relationships among A-genome species of the genus Oryza revealed by intron sequences of four nuclear genes. New Phytol. 167, 249–265 (2005). - PubMed
Publication types
MeSH terms
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

