Channel crossing: how are proteins shipped across the bacterial plasma membrane?
- PMID: 26370937
- PMCID: PMC4632601
- DOI: 10.1098/rstb.2015.0025
Channel crossing: how are proteins shipped across the bacterial plasma membrane?
Abstract
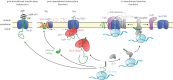
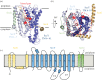
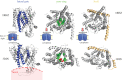
The structure of the first protein-conducting channel was determined more than a decade ago. Today, we are still puzzled by the outstanding problem of protein translocation--the dynamic mechanism underlying the consignment of proteins across and into membranes. This review is an attempt to summarize and understand the energy transducing capabilities of protein-translocating machines, with emphasis on bacterial systems: how polypeptides make headway against the lipid bilayer and how the process is coupled to the free energy associated with ATP hydrolysis and the transmembrane protein motive force. In order to explore how cargo is driven across the membrane, the known structures of the protein-translocation machines are set out against the background of the historic literature, and in the light of experiments conducted in their wake. The paper will focus on the bacterial general secretory (Sec) pathway (SecY-complex), and its eukaryotic counterpart (Sec61-complex), which ferry proteins across the membrane in an unfolded state, as well as the unrelated Tat system that assembles bespoke channels for the export of folded proteins.
Keywords: SecYEG; Tat; protein secretion; protein translocation.
© 2015 The Authors.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

