Kagami-Ogata syndrome: a clinically recognizable upd(14)pat and related disorder affecting the chromosome 14q32.2 imprinted region
- PMID: 26377239
- PMCID: PMC4771937
- DOI: 10.1038/jhg.2015.113
Kagami-Ogata syndrome: a clinically recognizable upd(14)pat and related disorder affecting the chromosome 14q32.2 imprinted region
Abstract
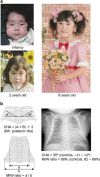
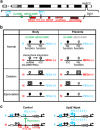
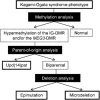
Human chromosome 14q32.2 carries paternally expressed genes including DLK1 and RTL1, and maternally expressed genes including MEG3 and RTL1as, along with the germline-derived DLK1-MEG3 intergenic differentially methylated region (IG-DMR) and the postfertilization-derived MEG3-DMR. Consistent with this, paternal uniparental disomy 14 (upd(14)pat), and epimutations (hypermethylations) and microdeletions affecting the IG-DMR and/or the MEG3-DMR of maternal origin, result in a unique phenotype associated with characteristic face, a small bell-shaped thorax with coat-hanger appearance of the ribs, abdominal wall defects, placentomegaly and polyhydramnios. Recently, the name 'Kagami-Ogata syndrome' (KOS) has been approved for this clinically recognizable disorder. Here, we review the current knowledge about KOS. Important findings include the following: (1) the facial 'gestalt' and the increased coat-hanger angle constitute pathognomonic features from infancy through childhood/puberty; (2) the unmethylated IG-DMR and MEG3-DMR of maternal origin function as the imprinting control centers in the placenta and body respectively, with a hierarchical interaction regulated by the IG-DMR for the methylation pattern of the MEG3-DMR in the body; (3) RTL1 expression level becomes ~2.5 times increased in the absence of functional RTL1as-encoded microRNAs that act as a trans-acting repressor for RTL1; (4) excessive RTL1 expression and absent MEG expression constitute the primary underlying factor for the phenotypic development; and (5) upd(14)pat accounts for approximately two-thirds of KOS patients, and epimutations and microdeletions are identified with a similar frequency. Furthermore, we refer to diagnostic and therapeutic implications.
Figures


Similar articles
-
Paternal uniparental disomy chromosome 14-like syndrome due a maternal de novo 160 kb deletion at the 14q32.2 region not encompassing the IG- and the MEG3-DMRs: Patient report and genotype-phenotype correlation.Am J Med Genet A. 2015 Dec;167A(12):3130-8. doi: 10.1002/ajmg.a.37293. Epub 2015 Sep 3. Am J Med Genet A. 2015. PMID: 26333487
-
Two infants with mild, atypical clinical features of Kagami-Ogata syndrome caused by epimutation.Eur J Med Genet. 2022 Oct;65(10):104580. doi: 10.1016/j.ejmg.2022.104580. Epub 2022 Aug 8. Eur J Med Genet. 2022. PMID: 35953028
-
Kagami-Ogata Syndrome.2024 Oct 24. In: Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Amemiya A, editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993–2025. 2024 Oct 24. In: Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Amemiya A, editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993–2025. PMID: 39446997 Free Books & Documents. Review.
-
New insights into the imprinted MEG8-DMR in 14q32 and clinical and molecular description of novel patients with Temple syndrome.Eur J Hum Genet. 2017 Aug;25(8):935-945. doi: 10.1038/ejhg.2017.91. Epub 2017 Jun 21. Eur J Hum Genet. 2017. PMID: 28635951 Free PMC article.
-
Molecular mechanisms regulating phenotypic outcome in paternal and maternal uniparental disomy for chromosome 14.Epigenetics. 2008 Jul-Aug;3(4):181-7. doi: 10.4161/epi.3.4.6550. Epub 2008 Jul 2. Epigenetics. 2008. PMID: 18698157 Review.
Cited by
-
The role of eutherian-specific RTL1 in the nervous system and its implications for the Kagami-Ogata and Temple syndromes.Genes Cells. 2021 Mar;26(3):165-179. doi: 10.1111/gtc.12830. Epub 2021 Feb 16. Genes Cells. 2021. PMID: 33484574 Free PMC article.
-
The 14q32 maternally imprinted locus is a major source of longitudinally stable circulating microRNAs as measured by small RNA sequencing.Sci Rep. 2019 Oct 31;9(1):15787. doi: 10.1038/s41598-019-51948-6. Sci Rep. 2019. PMID: 31673048 Free PMC article. Clinical Trial.
-
Kagami-Ogata Syndrome: Case Series and Review of Literature.AJP Rep. 2021 Mar;11(2):e65-e75. doi: 10.1055/s-0041-1727287. Epub 2021 May 27. AJP Rep. 2021. PMID: 34055463 Free PMC article.
-
Quantitative assessment of coat-hanger ribs detected on three-dimensional ultrasound for prenatal diagnosis of Kagami-Ogata syndrome.J Obstet Gynaecol Res. 2022 Dec;48(12):3314-3318. doi: 10.1111/jog.15425. Epub 2022 Sep 10. J Obstet Gynaecol Res. 2022. PMID: 36087043 Free PMC article.
-
The origin of imprinting defects in Temple syndrome and comparison with other imprinting disorders.Epigenetics. 2018;13(8):822-828. doi: 10.1080/15592294.2018.1514233. Epub 2018 Sep 19. Epigenetics. 2018. PMID: 30227764 Free PMC article.
References
-
- da Rocha, S. T., Edwards, C. A., Ito, M., Ogata, T. & Ferguson-Smith, A. C. Genomic imprinting at the mammalian Dlk1-Dio3 domain. Trends Genet. 24, 306–316 (2008). - PubMed
-
- Kagami, M., Sekita, Y., Nishimura, G., Irie, M., Kato, F., Okada, M. et al. Deletions and epimutations affecting the human 14q32.2 imprinted region in individuals with paternal and maternal upd(14)-like phenotypes. Nat. Genet. 40, 237–242 (2008). - PubMed
-
- Hoffmann, K. & Heller, R. Uniparental disomies 7 and 14. Best Pract. Res. Clin. Endocrinol. Metab. 25, 77–100 (2011). - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
