Independent Mechanisms Target SMCHD1 to Trimethylated Histone H3 Lysine 9-Modified Chromatin and the Inactive X Chromosome
- PMID: 26391951
- PMCID: PMC4628070
- DOI: 10.1128/MCB.00432-15
Independent Mechanisms Target SMCHD1 to Trimethylated Histone H3 Lysine 9-Modified Chromatin and the Inactive X Chromosome
Abstract
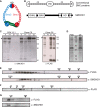
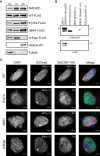
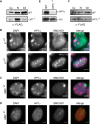
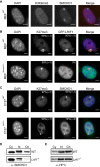
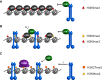
The chromosomal protein SMCHD1 plays an important role in epigenetic silencing at diverse loci, including the inactive X chromosome, imprinted genes, and the facioscapulohumeral muscular dystrophy locus. Although homology with canonical SMC family proteins suggests a role in chromosome organization, the mechanisms underlying SMCHD1 function and target site selection remain poorly understood. Here we show that SMCHD1 forms an active GHKL-ATPase homodimer, contrasting with canonical SMC complexes, which exist as tripartite ring structures. Electron microscopy analysis demonstrates that SMCHD1 homodimers structurally resemble prokaryotic condensins. We further show that the principal mechanism for chromatin loading of SMCHD1 involves an LRIF1-mediated interaction with HP1γ at trimethylated histone H3 lysine 9 (H3K9me3)-modified chromatin sites on the chromosome arms. A parallel pathway accounts for chromatin loading at a minority of sites, notably the inactive X chromosome. Together, our results provide key insights into SMCHD1 function and target site selection.
Copyright © 2015 Brideau et al.
Figures




References
-
- Ashe A, Morgan DK, Whitelaw NC, Bruxner TJ, Vickaryous NK, Cox LL, Butterfield NC, Wicking C, Blewitt ME, Wilkins SJ, Anderson GJ, Cox TC, Whitelaw E. 2008. A genome-wide screen for modifiers of transgene variegation identifies genes with critical roles in development. Genome Biol 9:R182. doi:10.1186/gb-2008-9-12-r182. - DOI - PMC - PubMed
-
- Blewitt ME, Gendrel AV, Pang Z, Sparrow DB, Whitelaw N, Craig JM, Apedaile A, Hilton DJ, Dunwoodie SL, Brockdorff N, Kay GF, Whitelaw E. 2008. SmcHD1, containing a structural-maintenance-of-chromosomes hinge domain, has a critical role in X inactivation. Nat Genet 40:663–669. doi:10.1038/ng.142. - DOI - PubMed
-
- Mould AW, Pang Z, Pakusch M, Tonks ID, Stark M, Carrie D, Mukhopadhyay P, Seidel A, Ellis JJ, Deakin J, Wakefield MJ, Krause L, Blewitt ME, Kay GF. 2013. Smchd1 regulates a subset of autosomal genes subject to monoallelic expression in addition to being critical for X inactivation. Epigenetics Chromatin 6:19. doi:10.1186/1756-8935-6-19. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
