Analytical Ultracentrifugation as a Tool to Study Nonspecific Protein-DNA Interactions
- PMID: 26412658
- PMCID: PMC5009906
- DOI: 10.1016/bs.mie.2015.04.009
Analytical Ultracentrifugation as a Tool to Study Nonspecific Protein-DNA Interactions
Abstract
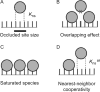
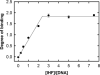
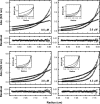
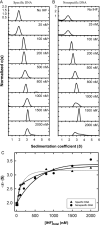
Analytical ultracentrifugation (AUC) is a powerful tool that can provide thermodynamic information on associating systems. Here, we discuss how to use the two fundamental AUC applications, sedimentation velocity (SV), and sedimentation equilibrium (SE), to study nonspecific protein-nucleic acid interactions, with a special emphasis on how to analyze the experimental data to extract thermodynamic information. We discuss three specific applications of this approach: (i) determination of nonspecific binding stoichiometry of E. coli integration host factor protein to dsDNA, (ii) characterization of nonspecific binding properties of Adenoviral IVa2 protein to dsDNA using SE-AUC, and (iii) analysis of the competition between specific and nonspecific DNA-binding interactions observed for E. coli integration host factor protein assembly on dsDNA. These approaches provide powerful tools that allow thermodynamic interrogation and thus a mechanistic understanding of how proteins bind nucleic acids by both specific and nonspecific interactions.
Keywords: Nonspecific binding; Specific and nonspecific competitive binding.
© 2015 Elsevier Inc. All rights reserved.
Figures
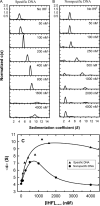

References
-
- Aeling KA, Opel ML, Steffen NR, Tretyachenko-Ladokhina V, Hatfield GW, Lathrop RH, et al. Indirect recognition in sequence-specific DNA binding by Escherichia coli integration host factor: The role of DNA deformation energy. The Journal of Biological Chemistry. 2006;281(51):39236–39248. - PubMed
-
- Bujalowski W. Thermodynamic and kinetic methods of analyses of protein-nucleic acid interactions. From simpler to more complex systems. Chemical Reviews. 2006;106(2):556–606. - PubMed
-
- Bujalowski W, Lohman TM, Anderson CF. On the cooperative binding of large ligands to a one-dimensional homogeneous lattice: The generalized three-state lattice model. Biopolymers. 1989;28(9):1637–1643. - PubMed
-
- Bujalowski W, Overman LB, Lohman TM. Binding mode transitions of Escherichia coli single strand binding protein-single-stranded DNA complexes. Cation, anion, pH, and binding density effects. The Journal of Biological Chemistry. 1988;263(10):4629–4640. - PubMed
-
- Bustamante C, Bryant Z, Smith SB. Ten years of tension: Single-molecule DNA mechanics. Nature. 2003;421(6921):423–427. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

