Deep sequencing analysis of transcriptomes in Aspergillus flavus in response to resveratrol
- PMID: 26420172
- PMCID: PMC4589122
- DOI: 10.1186/s12866-015-0513-6
Deep sequencing analysis of transcriptomes in Aspergillus flavus in response to resveratrol
Abstract
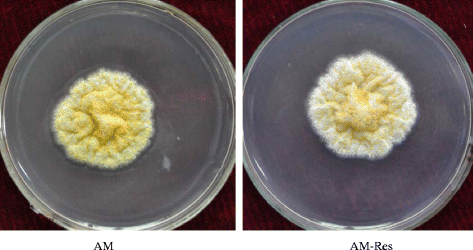
Background: Resveratrol has been reported as a natural phytoalexin that inhibits infection or the growth of certain fungi including Aspergillus flavus. Our previous research revealed that aflatoxin production in A. flavus was reduced in medium with resveratrol. To understand the molecular mechanism of the A. flavus response to resveratrol treatment, the high-throughput paired-end RNA-Seq was applied to analyze the transcriptomic profiles of A. flavus.
Results: In total, 366 and 87 genes of A. flavus were significantly up- and down- regulated, respectively, when the fungus was treated with resveratrol. Gene Ontology (GO) functional enrichment analysis revealed that 48 significantly differentially expressed genes were involved in 6 different terms. Most genes in the aflatoxin biosynthetic pathway genes cluster (#54) did not show a significant change when A. flavus was treated with resveratrol, but 23 of the 30 genes in the #54 cluster were down-regulated. The transcription of aflA and aflB was significantly suppressed under resveratrol treatment, resulting in an insufficient amount of the starter unit hexanoate for aflatoxin biosynthesis. In addition, resveratrol significantly increased the activity of antioxidative enzymes that destroy radicals, leading to decreased aflatoxin production. Moreover, stuA, fluG, flbC, and others genes involved in mycelial and conidial development were down-regulated, which disrupted the cell's orderly differentiation and blocked conidia formation and mycelia development. The transcripts of laeA and veA were slightly inhibited by resveratrol, which may partly decrease aflatoxin production and depress conidia formation.
Conclusions: Resveratrol can affect the expression of A. flavus genes that are related to developmental and secondary metabolic processes, resulting in decreased aflatoxin production and conidia formation and could also cause abnormal mycelia development. These results provide insight into the transcriptome of A. flavus in response to resveratrol and a new clew for further study in regulation of aflatoxin biosynthesis in A. flavus.
Figures



References
-
- Williams JH, Phillips TD, Jolly PE, Stiles JK, Jolly CM, Aggarwal D. Human aflatoxicosis in developing countries: a review of toxicology, exposure, potential health consequences, and interventions. Am J Clin Nutrit. 2004;80(5):1106–1122. - PubMed
-
- Burow GB, Nesbitt TC, Dunlap J, Keller NP. Seed lipoxygenase products modulate Aspergillus mycotoxin biosynthesis. Mol. Plant Microbe Interact. 1997;10(3):380–387. doi: 10.1094/MPMI.1997.10.3.380. - DOI
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources

