Characterization of Bacterial, Archaeal and Eukaryote Symbionts from Antarctic Sponges Reveals a High Diversity at a Three-Domain Level and a Particular Signature for This Ecosystem
- PMID: 26421612
- PMCID: PMC4589366
- DOI: 10.1371/journal.pone.0138837
Characterization of Bacterial, Archaeal and Eukaryote Symbionts from Antarctic Sponges Reveals a High Diversity at a Three-Domain Level and a Particular Signature for This Ecosystem
Abstract
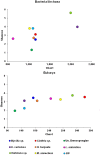
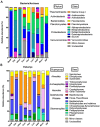
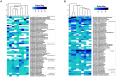
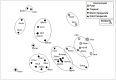
Sponge-associated microbial communities include members from the three domains of life. In the case of bacteria, they are diverse, host specific and different from the surrounding seawater. However, little is known about the diversity and specificity of Eukarya and Archaea living in association with marine sponges. This knowledge gap is even greater regarding sponges from regions other than temperate and tropical environments. In Antarctica, marine sponges are abundant and important members of the benthos, structuring the Antarctic marine ecosystem. In this study, we used high throughput ribosomal gene sequencing to investigate the three-domain diversity and community composition from eight different Antarctic sponges. Taxonomic identification reveals that they belong to families Acarnidae, Chalinidae, Hymedesmiidae, Hymeniacidonidae, Leucettidae, Microcionidae, and Myxillidae. Our study indicates that there are different diversity and similarity patterns between bacterial/archaeal and eukaryote microbial symbionts from these Antarctic marine sponges, indicating inherent differences in how organisms from different domains establish symbiotic relationships. In general, when considering diversity indices and number of phyla detected, sponge-associated communities are more diverse than the planktonic communities. We conclude that three-domain microbial communities from Antarctic sponges are different from surrounding planktonic communities, expanding previous observations for Bacteria and including the Antarctic environment. Furthermore, we reveal differences in the composition of the sponge associated bacterial assemblages between Antarctic and tropical-temperate environments and the presence of a highly complex microbial eukaryote community, suggesting a particular signature for Antarctic sponges, different to that reported from other ecosystems.
Conflict of interest statement
Figures
Similar articles
-
Diverse microbial communities inhabit Antarctic sponges.Environ Microbiol. 2004 Mar;6(3):288-300. doi: 10.1111/j.1462-2920.2004.00570.x. Environ Microbiol. 2004. PMID: 14871212
-
The Microbial Ecology of Antarctic Sponges.Microb Ecol. 2025 May 17;88(1):44. doi: 10.1007/s00248-025-02543-y. Microb Ecol. 2025. PMID: 40382475 Free PMC article. Review.
-
Dynamic microbiome diversity shaping the adaptation of sponge holobionts in coastal waters.Microbiol Spectr. 2024 Nov 5;12(11):e0144824. doi: 10.1128/spectrum.01448-24. Epub 2024 Oct 14. Microbiol Spectr. 2024. PMID: 39400157 Free PMC article.
-
Unveiling microbial guilds and symbiotic relationships in Antarctic sponge microbiomes.Sci Rep. 2024 Mar 16;14(1):6371. doi: 10.1038/s41598-024-56480-w. Sci Rep. 2024. PMID: 38493232 Free PMC article.
-
Diversity and biotechnological potential of microorganisms associated with marine sponges.Appl Microbiol Biotechnol. 2014 Sep;98(17):7331-47. doi: 10.1007/s00253-014-5861-x. Epub 2014 Jul 10. Appl Microbiol Biotechnol. 2014. PMID: 25005058 Review.
Cited by
-
Inhabitancy of active Nitrosopumilus-like ammonia-oxidizing archaea and Nitrospira nitrite-oxidizing bacteria in the sponge Theonella swinhoei.Sci Rep. 2016 Apr 26;6:24966. doi: 10.1038/srep24966. Sci Rep. 2016. PMID: 27113140 Free PMC article.
-
The Sponge-Associated Fungus Eurotium chevalieri MUT 2316 and its Bioactive Molecules: Potential Applications in the Field of Antifouling.Mar Biotechnol (NY). 2019 Dec;21(6):743-752. doi: 10.1007/s10126-019-09920-y. Epub 2019 Sep 7. Mar Biotechnol (NY). 2019. PMID: 31494811
-
Antarctic sponges from the Terra Nova Bay (Ross Sea) host a diversified bacterial community.Sci Rep. 2019 Nov 6;9(1):16135. doi: 10.1038/s41598-019-52491-0. Sci Rep. 2019. PMID: 31695084 Free PMC article.
-
A Metataxonomic Approach Reveals Diversified Bacterial Communities in Antarctic Sponges.Mar Drugs. 2021 Mar 22;19(3):173. doi: 10.3390/md19030173. Mar Drugs. 2021. PMID: 33810171 Free PMC article.
-
First Molecular Identification of Symbiotic Archaea in a Sponge Collected from the Persian Gulf, Iran.Open Microbiol J. 2018 Oct 17;12:323-332. doi: 10.2174/1874285801812010323. eCollection 2018. Open Microbiol J. 2018. PMID: 30450139 Free PMC article.
References
-
- Margulis L, Fester R (Eds.) Symbiosis as a source of evolutionary innovation: speciation and morphogenesis. MIT Press; 1991. - PubMed
-
- Gilbert SF. Symbiosis as the way of eukaryotic life: The dependent co-origination of the body. J Biosci. 2014;39(2): 201–209. - PubMed
-
- Vacelet J. Electron-microscope study of association between bacteria and sponges of genus Verongia (Dictyoceratida). J Microsc Biol Cell. 1975;23(3): 271.
-
- Hentschel U, Usher KM, Taylor MW. Marine sponges as microbial fermenters. FEMS Microbiol Ecol. 2006;55(2): 167–177. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

