A Bayesian approach for inducing sparsity in generalized linear models with multi-category response
- PMID: 26423345
- PMCID: PMC4597416
- DOI: 10.1186/1471-2105-16-S13-S13
A Bayesian approach for inducing sparsity in generalized linear models with multi-category response
Abstract
Background: The dimension and complexity of high-throughput gene expression data create many challenges for downstream analysis. Several approaches exist to reduce the number of variables with respect to small sample sizes. In this study, we utilized the Generalized Double Pareto (GDP) prior to induce sparsity in a Bayesian Generalized Linear Model (GLM) setting. The approach was evaluated using a publicly available microarray dataset containing 99 samples corresponding to four different prostate cancer subtypes.
Results: A hierarchical Sparse Bayesian GLM using GDP prior (SBGG) was developed to take into account the progressive nature of the response variable. We obtained an average overall classification accuracy between 82.5% and 94%, which was higher than Support Vector Machine, Random Forest or a Sparse Bayesian GLM using double exponential priors. Additionally, SBGG outperforms the other 3 methods in correctly identifying pre-metastatic stages of cancer progression, which can prove extremely valuable for therapeutic and diagnostic purposes. Importantly, using Geneset Cohesion Analysis Tool, we found that the top 100 genes produced by SBGG had an average functional cohesion p-value of 2.0E-4 compared to 0.007 to 0.131 produced by the other methods.
Conclusions: Using GDP in a Bayesian GLM model applied to cancer progression data results in better subclass prediction. In particular, the method identifies pre-metastatic stages of prostate cancer with substantially better accuracy and produces more functionally relevant gene sets.
Figures

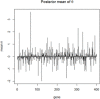

References
-
- Devore J, Peck R. Statistics: The Exploration and Analysis of Data. Duxbury, Pacific Grove CA; 1997.
-
- Pan W. A comparative review of statistical methods for discovering differentially expressed genes in replicated microarray experiments. Bioinformatics. 1996;18(4):546–554. - PubMed
-
- Dudoit S, Fridlyand J, Speed TP. Comparison of discrimination methods for the classification of tumors using gene expression data. J Am Stat Assoc. 2002;97(457):77–87. doi: 10.1198/016214502753479248. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical

