Quadruplex formation by both G-rich and C-rich DNA strands of the C9orf72 (GGGGCC)8•(GGCCCC)8 repeat: effect of CpG methylation
- PMID: 26432832
- PMCID: PMC4787773
- DOI: 10.1093/nar/gkv1008
Quadruplex formation by both G-rich and C-rich DNA strands of the C9orf72 (GGGGCC)8•(GGCCCC)8 repeat: effect of CpG methylation
Abstract
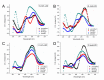
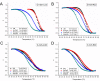
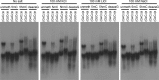
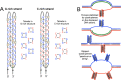
Unusual DNA/RNA structures of the C9orf72 repeat may participate in repeat expansions or pathogenesis of amyotrophic lateral sclerosis and frontotemporal dementia. Expanded repeats are CpG methylated with unknown consequences. Typically, quadruplex structures form by G-rich but not complementary C-rich strands. Using CD, UV and electrophoresis, we characterized the structures formed by (GGGGCC)8 and (GGCCCC)8 strands with and without 5-methylcytosine (5mCpG) or 5-hydroxymethylcytosine (5hmCpG) methylation. All strands formed heterogenous mixtures of structures, with features of quadruplexes (at pH 7.5, in K(+), Na(+) or Li(+)), but no feature typical of i-motifs. C-rich strands formed quadruplexes, likely stabilized by G•C•G•C-tetrads and C•C•C•C-tetrads. Unlike G•G•G•G-tetrads, some G•C•G•C-tetrad conformations do not require the N7-Guanine position, hence C9orf72 quadruplexes still formed when N7-deazaGuanine replace all Guanines. 5mCpG and 5hmCpG increased and decreased the thermal stability of these structures. hnRNPK, through band-shift analysis, bound C-rich but not G-rich strands, with a binding preference of unmethylated > 5hmCpG > 5mCpG, where methylated DNA-protein complexes were retained in the wells, distinct from unmethylated complexes. Our findings suggest that for C-rich sequences interspersed with G-residues, one must consider quadruplex formation and that methylation of quadruplexes may affect epigenetic processes.
© The Author(s) 2015. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

Similar articles
-
Thermodynamic and spectroscopic investigations of TMPyP4 association with guanine- and cytosine-rich DNA and RNA repeats of C9orf72.Biochem Biophys Res Commun. 2018 Jan 22;495(4):2410-2417. doi: 10.1016/j.bbrc.2017.12.108. Epub 2017 Dec 20. Biochem Biophys Res Commun. 2018. PMID: 29274339
-
Stress-induced acidification may contribute to formation of unusual structures in C9orf72-repeats.Biochim Biophys Acta Gen Subj. 2018 Jun;1862(6):1482-1491. doi: 10.1016/j.bbagen.2018.03.001. Epub 2018 Mar 15. Biochim Biophys Acta Gen Subj. 2018. PMID: 29550431
-
The disease-associated r(GGGGCC)n repeat from the C9orf72 gene forms tract length-dependent uni- and multimolecular RNA G-quadruplex structures.J Biol Chem. 2013 Apr 5;288(14):9860-9866. doi: 10.1074/jbc.C113.452532. Epub 2013 Feb 19. J Biol Chem. 2013. PMID: 23423380 Free PMC article.
-
Human telomere, oncogenic promoter and 5'-UTR G-quadruplexes: diverse higher order DNA and RNA targets for cancer therapeutics.Nucleic Acids Res. 2007;35(22):7429-55. doi: 10.1093/nar/gkm711. Epub 2007 Oct 2. Nucleic Acids Res. 2007. PMID: 17913750 Free PMC article. Review.
-
Electronic excitations in Guanine quadruplexes.Top Curr Chem. 2015;356:183-201. doi: 10.1007/128_2013_511. Top Curr Chem. 2015. PMID: 24563011 Review.
Cited by
-
C9orf72-FTD/ALS pathogenesis: evidence from human neuropathological studies.Acta Neuropathol. 2019 Jan;137(1):1-26. doi: 10.1007/s00401-018-1921-0. Epub 2018 Oct 27. Acta Neuropathol. 2019. PMID: 30368547 Free PMC article. Review.
-
G-quadruplex DNA: a novel target for drug design.Cell Mol Life Sci. 2021 Oct;78(19-20):6557-6583. doi: 10.1007/s00018-021-03921-8. Epub 2021 Aug 30. Cell Mol Life Sci. 2021. PMID: 34459951 Free PMC article. Review.
-
Close encounters: Moving along bumps, breaks, and bubbles on expanded trinucleotide tracts.DNA Repair (Amst). 2017 Aug;56:144-155. doi: 10.1016/j.dnarep.2017.06.017. Epub 2017 Jun 9. DNA Repair (Amst). 2017. PMID: 28690053 Free PMC article. Review.
-
Non-duplex G-Quadruplex DNA Structure: A Developing Story from Predicted Sequences to DNA Structure-Dependent Epigenetics and Beyond.Acc Chem Res. 2021 Jan 5;54(1):46-56. doi: 10.1021/acs.accounts.0c00431. Epub 2020 Dec 21. Acc Chem Res. 2021. PMID: 33347280 Free PMC article.
-
G-quadruplex structures formed by human telomeric DNA and C9orf72 hexanucleotide repeats.Biophys Rev. 2019 Jun;11(3):389-393. doi: 10.1007/s12551-019-00545-y. Epub 2019 May 24. Biophys Rev. 2019. PMID: 31127470 Free PMC article. No abstract available.
References
-
- Pearson C.E., Nichol Edamura K., Cleary J.D. Repeat instability: mechanisms of dynamic mutations. Nat. Rev. Genet. 2005;6:729–742. - PubMed
-
- Belzil V.V., Bauer P.O., Prudencio M., Gendron T.F., Stetler C.T., Yan I.K., Pregent L., Daughrity L., Baker M.C., Rademakers R., et al. Reduced C9orf72 gene expression in c9FTD/ALS is caused by histone trimethylation, an epigenetic event detectable in blood. Acta Neuropathol. 2013;126:895–905. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

