Temporal small RNA transcriptome profiling unraveled partitioned miRNA expression in developing maize endosperms between reciprocal crosses
- PMID: 26442057
- PMCID: PMC4584948
- DOI: 10.3389/fpls.2015.00744
Temporal small RNA transcriptome profiling unraveled partitioned miRNA expression in developing maize endosperms between reciprocal crosses
Abstract
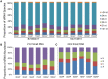
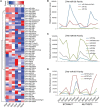
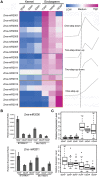
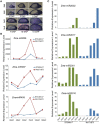
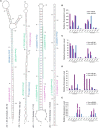
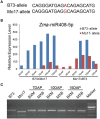
In angiosperms, the endosperm nurtures the embryo and provides nutrients for seed germination. To identify the expression pattern of small interfering RNA in the developing maize endosperm, we have performed high-throughput small RNA transcriptome sequencing of kernels at 0, 3, and 5 days after pollination (DAP) and endosperms at 7, 10, and 15 DAP using B73 and Mo17 reciprocal crosses in previous study. Here, we further explored these small RNA-seq data to investigate the potential roles of miRNAs in regulating the gene expression process. In total, 57 conserved miRNAs and 18 novel miRNAs were observed highly expressed in maize endosperm. Temporal expression profiling indicated that these miRNAs exhibited dynamic and partitioned expression patterns at different developmental stages between maize reciprocal crosses, and quantitative RT-PCR results further confirmed our observation. In addition, we found a subset of distinct tandem miRNAs are generated from a single stem-loop structure in maize that might be conserved in monocots. Furthermore, a SNP variation of Zma-miR408-5p at 11th base position was characterized between B73 and Mo17 which might lead to completely different functions in repressing targets. More interestingly, Zma-miR408-5p exhibited B73-biased expression pattern in the B73 and Mo17 reciprocal hybrid endosperms at 7, 10, and 15 DAP according to the reads abundance with SNPs and CAPS experiment. Together, this study suggests that miRNA plays a crucial role in regulating endosperm development, and exhibited distinct expression patterns in developing endosperm between maize reciprocal crosses.
Keywords: maize endosperm; miRNA profiling; partitioned expression; reciprocal cross; tandem miRNA.
Figures
Similar articles
-
Dynamic parent-of-origin effects on small interfering RNA expression in the developing maize endosperm.BMC Plant Biol. 2014 Jul 24;14:192. doi: 10.1186/s12870-014-0192-8. BMC Plant Biol. 2014. PMID: 25055833 Free PMC article.
-
Dynamic expression of imprinted genes associates with maternally controlled nutrient allocation during maize endosperm development.Plant Cell. 2013 Sep;25(9):3212-27. doi: 10.1105/tpc.113.115592. Epub 2013 Sep 20. Plant Cell. 2013. PMID: 24058158 Free PMC article.
-
High-Throughput Sequencing of Small RNA Transcriptomes in Maize Kernel Identifies miRNAs Involved in Embryo and Endosperm Development.Genes (Basel). 2017 Dec 14;8(12):385. doi: 10.3390/genes8120385. Genes (Basel). 2017. PMID: 29240690 Free PMC article.
-
The functions of the endosperm during seed germination.Plant Cell Physiol. 2014 Sep;55(9):1521-33. doi: 10.1093/pcp/pcu089. Epub 2014 Jun 24. Plant Cell Physiol. 2014. PMID: 24964910 Review.
-
Maize Endosperm Development: Tissues, Cells, Molecular Regulation and Grain Quality Improvement.Front Plant Sci. 2022 Mar 7;13:852082. doi: 10.3389/fpls.2022.852082. eCollection 2022. Front Plant Sci. 2022. PMID: 35330868 Free PMC article. Review.
Cited by
-
Integrated transcriptome, small RNA, and degradome analysis reveals the complex network regulating starch biosynthesis in maize.BMC Genomics. 2019 Jul 11;20(1):574. doi: 10.1186/s12864-019-5945-1. BMC Genomics. 2019. PMID: 31296166 Free PMC article.
-
MicroRNA transcriptomic analysis of the sixth leaf of maize (Zea mays L.) revealed a regulatory mechanism of jointing stage heterosis.BMC Plant Biol. 2020 Nov 30;20(1):541. doi: 10.1186/s12870-020-02751-3. BMC Plant Biol. 2020. PMID: 33256592 Free PMC article.
-
The pivotal role of small non-coding RNAs in the regulation of seed development.Plant Cell Rep. 2017 May;36(5):653-667. doi: 10.1007/s00299-017-2120-5. Epub 2017 Mar 13. Plant Cell Rep. 2017. PMID: 28289886 Review.
-
Development of Incompletely Fused Carpels in Maize Ovary Revealed by miRNA, Target Gene and Phytohormone Analysis.Front Plant Sci. 2017 Apr 3;8:463. doi: 10.3389/fpls.2017.00463. eCollection 2017. Front Plant Sci. 2017. PMID: 28421097 Free PMC article.
-
Heterotic patterns of primary and secondary metabolites in the oilseed crop Brassica juncea.Heredity (Edinb). 2019 Sep;123(3):318-336. doi: 10.1038/s41437-019-0213-3. Epub 2019 Mar 25. Heredity (Edinb). 2019. PMID: 30911141 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

