Tracing the Spread of Clostridium difficile Ribotype 027 in Germany Based on Bacterial Genome Sequences
- PMID: 26444881
- PMCID: PMC4596877
- DOI: 10.1371/journal.pone.0139811
Tracing the Spread of Clostridium difficile Ribotype 027 in Germany Based on Bacterial Genome Sequences
Abstract
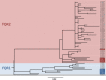
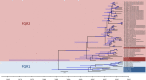
We applied whole-genome sequencing to reconstruct the spatial and temporal dynamics underpinning the expansion of Clostridium difficile ribotype 027 in Germany. Based on re-sequencing of genomes from 57 clinical C. difficile isolates, which had been collected from hospitalized patients at 36 locations throughout Germany between 1990 and 2012, we demonstrate that C. difficile genomes have accumulated sequence variation sufficiently fast to document the pathogen's spread at a regional scale. We detected both previously described lineages of fluoroquinolone-resistant C. difficile ribotype 027, FQR1 and FQR2. Using Bayesian phylogeographic analyses, we show that fluoroquinolone-resistant C. difficile 027 was imported into Germany at least four times, that it had been widely disseminated across multiple federal states even before the first outbreak was noted in 2007, and that it has continued to spread since.
Conflict of interest statement
Figures

Similar articles
-
Antibiotic profiling of Clostridium difficile ribotype 176--A multidrug resistant relative to C. difficile ribotype 027.Anaerobe. 2015 Dec;36:88-90. doi: 10.1016/j.anaerobe.2015.07.009. Epub 2015 Aug 6. Anaerobe. 2015. PMID: 26256807
-
[Epidemiological study of Clostridium difficile strains isolated in Jean-Verdier-René-Muret hospitals from 2001 to 2007].Pathol Biol (Paris). 2008 Nov-Dec;56(7-8):412-6. doi: 10.1016/j.patbio.2008.07.009. Epub 2008 Oct 7. Pathol Biol (Paris). 2008. PMID: 18842360 French.
-
Imipenem Resistance in Clostridium difficile Ribotype 017, Portugal.Emerg Infect Dis. 2018 Apr;24(4):741-745. doi: 10.3201/eid2404.170095. Emerg Infect Dis. 2018. PMID: 29553322 Free PMC article.
-
Reprint of New opportunities for improved ribotyping of C. difficile clinical isolates by exploring their genomes.J Microbiol Methods. 2013 Dec;95(3):425-40. doi: 10.1016/j.mimet.2013.09.009. Epub 2013 Sep 16. J Microbiol Methods. 2013. PMID: 24050948 Review.
-
New opportunities for improved ribotyping of C. difficile clinical isolates by exploring their genomes.J Microbiol Methods. 2013 Jun;93(3):257-72. doi: 10.1016/j.mimet.2013.02.013. Epub 2013 Mar 29. J Microbiol Methods. 2013. PMID: 23545446 Review.
Cited by
-
Advances in the Microbiome: Applications to Clostridium difficile Infection.J Clin Med. 2016 Sep 21;5(9):83. doi: 10.3390/jcm5090083. J Clin Med. 2016. PMID: 27657145 Free PMC article. Review.
-
Scarless deletion of up to seven methyl-accepting chemotaxis genes with an optimized method highlights key function of CheM in Salmonella Typhimurium.PLoS One. 2017 Feb 17;12(2):e0172630. doi: 10.1371/journal.pone.0172630. eCollection 2017. PLoS One. 2017. PMID: 28212413 Free PMC article.
-
The largely unnoticed spread of Clostridioides difficile PCR ribotype 027 in Germany after 2010.Infect Prev Pract. 2020 Nov 3;2(4):100102. doi: 10.1016/j.infpip.2020.100102. eCollection 2020 Dec. Infect Prev Pract. 2020. PMID: 34368730 Free PMC article. Review.
-
CRISPR Diversity and Microevolution in Clostridium difficile.Genome Biol Evol. 2016 Sep 19;8(9):2841-55. doi: 10.1093/gbe/evw203. Genome Biol Evol. 2016. PMID: 27576538 Free PMC article.
-
Convergent Loss of ABC Transporter Genes From Clostridioides difficile Genomes Is Associated With Impaired Tyrosine Uptake and p-Cresol Production.Front Microbiol. 2018 May 8;9:901. doi: 10.3389/fmicb.2018.00901. eCollection 2018. Front Microbiol. 2018. PMID: 29867812 Free PMC article.
References
-
- Suetens C, Hopkins S, Kolman J, Högberg LD. Point prevalence survey of healthcareassociated infections and antimicrobial use in European acute care hospitals 2011–2012: European Centre for Disease Prevention and Control; 2013. Available from: www.ecdc.europa.eu.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

