Linking Biosynthetic Gene Clusters to their Metabolites via Pathway- Targeted Molecular Networking
- PMID: 26456470
- PMCID: PMC5055756
- DOI: 10.2174/1568026616666151012111046
Linking Biosynthetic Gene Clusters to their Metabolites via Pathway- Targeted Molecular Networking
Abstract
The connection of microbial biosynthetic gene clusters to the small molecule metabolites they encode is central to the discovery and characterization of new metabolic pathways with ecological and pharmacological potential. With increasing microbial genome sequence information being deposited into publicly available databases, it is clear that microbes have the coding capacity for many more biologically active small molecules than previously realized. Of increasing interest are the small molecules encoded by the human microbiome, as these metabolites likely mediate a variety of currently uncharacterized human-microbe interactions that influence health and disease. In this mini-review, we describe the ongoing biosynthetic, structural, and functional characterizations of the genotoxic colibactin pathway in gut bacteria as a thematic example of linking biosynthetic gene clusters to their metabolites. We also highlight other natural products that are produced through analogous biosynthetic logic and comment on some current disconnects between bioinformatics predictions and experimental structural characterizations. Lastly, we describe the use of pathway-targeted molecular networking as a tool to characterize secondary metabolic pathways within complex metabolomes and to aid in downstream metabolite structural elucidation efforts.
Conflict of interest statement
The author(s) confirm that this article content has no conflicts of interest.
Figures
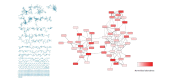

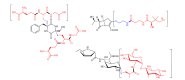
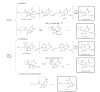
Similar articles
-
Recent development of antiSMASH and other computational approaches to mine secondary metabolite biosynthetic gene clusters.Brief Bioinform. 2019 Jul 19;20(4):1103-1113. doi: 10.1093/bib/bbx146. Brief Bioinform. 2019. PMID: 29112695 Free PMC article.
-
Bioinformatics assisted construction of the link between biosynthetic gene clusters and secondary metabolites in fungi.Biotechnol Adv. 2025 Jul-Aug;81:108547. doi: 10.1016/j.biotechadv.2025.108547. Epub 2025 Feb 28. Biotechnol Adv. 2025. PMID: 40024584 Review.
-
Identification of the Bacterial Biosynthetic Gene Clusters of the Oral Microbiome Illuminates the Unexplored Social Language of Bacteria during Health and Disease.mBio. 2019 Apr 16;10(2):e00321-19. doi: 10.1128/mBio.00321-19. mBio. 2019. PMID: 30992349 Free PMC article.
-
A Multi-Label Learning Framework for Predicting Chemical Classes and Biological Activities of Natural Products from Biosynthetic Gene Clusters.J Chem Ecol. 2023 Dec;49(11-12):681-695. doi: 10.1007/s10886-023-01452-z. Epub 2023 Oct 2. J Chem Ecol. 2023. PMID: 37779180
-
Studies on biosynthetic enzymes leading to structural and functional diversity of microbial natural products.Biosci Biotechnol Biochem. 2023 Jul 24;87(8):797-808. doi: 10.1093/bbb/zbad064. Biosci Biotechnol Biochem. 2023. PMID: 37226538 Review.
Cited by
-
Depurination of Colibactin-Derived Interstrand Cross-Links.Biochemistry. 2020 Feb 25;59(7):892-900. doi: 10.1021/acs.biochem.9b01070. Epub 2020 Feb 13. Biochemistry. 2020. PMID: 31977191 Free PMC article.
-
Structure elucidation of colibactin and its DNA cross-links.Science. 2019 Sep 6;365(6457):eaax2685. doi: 10.1126/science.aax2685. Epub 2019 Aug 8. Science. 2019. PMID: 31395743 Free PMC article.
-
Employing chemical synthesis to study the structure and function of colibactin, a "dark matter" metabolite.Nat Prod Rep. 2020 Nov 18;37(11):1532-1548. doi: 10.1039/d0np00072h. Nat Prod Rep. 2020. PMID: 33174565 Free PMC article. Review.
-
Marine Mammal Microbiota Yields Novel Antibiotic with Potent Activity Against Clostridium difficile.ACS Infect Dis. 2018 Jan 12;4(1):59-67. doi: 10.1021/acsinfecdis.7b00105. Epub 2017 Oct 18. ACS Infect Dis. 2018. PMID: 29043783 Free PMC article.
-
Comparative mass spectrometry-based metabolomics strategies for the investigation of microbial secondary metabolites.Nat Prod Rep. 2017 Jan 4;34(1):6-24. doi: 10.1039/c6np00048g. Nat Prod Rep. 2017. PMID: 27604382 Free PMC article. Review.
References
-
- Butler M.S., Robertson A.A., Cooper M.A. Natural product and natural product derived drugs in clinical trials. Nat. Prod. Rep. 2014;31(11):1612–1661. - PubMed
-
- Li J.W, Vederas J.C. Drug Discovery and Natural Products, End of an Era or an Endless Frontier? Science. 2009;325:161–165. - PubMed
-
- Demain A.L. Importance of microbial natural products and the need to revitalize their discovery. J. Ind. Microbiol. Biotechnol. 2014;41(2):185–201. - PubMed
-
- Challis G.L. Mining microbial genomes for new natural products and biosynthetic pathways. Microbiology. 2008;154(Pt 6):1555–1569. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
