Viral variants that initiate and drive maturation of V1V2-directed HIV-1 broadly neutralizing antibodies
- PMID: 26457756
- PMCID: PMC4637988
- DOI: 10.1038/nm.3963
Viral variants that initiate and drive maturation of V1V2-directed HIV-1 broadly neutralizing antibodies
Abstract
The elicitation of broadly neutralizing antibodies (bNAbs) is likely to be essential for a preventative HIV-1 vaccine, but this has not yet been achieved by immunization. In contrast, some HIV-1-infected individuals naturally mount bNAb responses during chronic infection, suggesting that years of maturation may be required for neutralization breadth. Recent studies have shown that viral diversification precedes the emergence of bNAbs, but the significance of this observation is unknown. Here we delineate the key viral events that drove neutralization breadth within the CAP256-VRC26 family of 33 monoclonal antibodies (mAbs) isolated from a superinfected individual. First, we identified minority viral variants, termed bNAb-initiating envelopes, that were distinct from both of the transmitted/founder (T/F) viruses and that efficiently engaged the bNAb precursor. Second, deep sequencing revealed a pool of diverse epitope variants (immunotypes) that were preferentially neutralized by broader members of the antibody lineage. In contrast, a 'dead-end' antibody sublineage unable to neutralize these immunotypes showed limited evolution and failed to develop breadth. Thus, early viral escape at key antibody-virus contact sites selects for antibody sublineages that can tolerate these changes, thereby providing a mechanism for the generation of neutralization breadth within a developing antibody lineage.
Figures
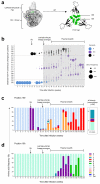



References
METHODS-ONLY REFERENCES
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

