Bcl6 middle domain repressor function is required for T follicular helper cell differentiation and utilizes the corepressor MTA3
- PMID: 26460037
- PMCID: PMC4629362
- DOI: 10.1073/pnas.1507312112
Bcl6 middle domain repressor function is required for T follicular helper cell differentiation and utilizes the corepressor MTA3
Abstract
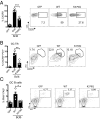
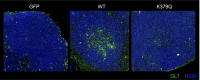
T follicular helper (Tfh) cells are essential providers of help to B cells. The transcription factor B-cell CLL/lymphoma 6 (Bcl6) is a lineage-defining regulator of Tfh cells and germinal center B cells. In B cells, Bcl6 has the potential to recruit distinct transcriptional corepressors through its BTB domain or its poorly characterized middle domain (also known as RDII), but in Tfh cells the roles of the Bcl6 middle domain have yet to be clarified. Mimicked acetylation of the Bcl6 middle domain (K379Q) in CD4 T cells results in significant reductions in Tfh differentiation in vivo. Blimp1 (Prdm1) is a potent inhibitor of Tfh cell differentiation. Although Bcl6 K379Q still bound to the Prdm1 cis-regulatory elements in Tfh cells, Prdm1 expression was derepressed. This was a result of the failure of Bcl6 K379Q to recruit metastasis-associated protein 3 (MTA3). The loss of Bcl6 function in Bcl6 K379Q-expressing CD4 T cells could be partially rescued by abrogating Prdm1 expression. In addition to Prdm1, we found that Bcl6 recruits MTA3 to multiple genes involved in Tfh cell biology, including genes important for cell migration, cell survival, and alternative differentiation pathways. Thus, Bcl6 middle domain mediated repression is a major mechanism of action by which Bcl6 controls CD4 T-cell fate and function.
Keywords: B-cell help; T follicular helper cells; germinal centers.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





Similar articles
-
Cutting edge: T follicular helper cell differentiation is defective in the absence of Bcl6 BTB repressor domain function.J Immunol. 2015 Jun 15;194(12):5599-603. doi: 10.4049/jimmunol.1500200. Epub 2015 May 8. J Immunol. 2015. PMID: 25957170 Free PMC article.
-
BCL6 programs lymphoma cells for survival and differentiation through distinct biochemical mechanisms.Blood. 2007 Sep 15;110(6):2067-74. doi: 10.1182/blood-2007-01-069575. Epub 2007 Jun 1. Blood. 2007. PMID: 17545502 Free PMC article.
-
An Inhibitory Role for the Transcription Factor Stat3 in Controlling IL-4 and Bcl6 Expression in Follicular Helper T Cells.J Immunol. 2015 Sep 1;195(5):2080-9. doi: 10.4049/jimmunol.1500335. Epub 2015 Jul 17. J Immunol. 2015. PMID: 26188063 Free PMC article.
-
Bcl6-Mediated Transcriptional Regulation of Follicular Helper T cells (TFH).Trends Immunol. 2021 Apr;42(4):336-349. doi: 10.1016/j.it.2021.02.002. Epub 2021 Mar 1. Trends Immunol. 2021. PMID: 33663954 Free PMC article. Review.
-
Differentiation of germinal center B cells and follicular helper T cells as viewed by tracking Bcl6 expression dynamics.Immunol Rev. 2012 May;247(1):120-32. doi: 10.1111/j.1600-065X.2012.01120.x. Immunol Rev. 2012. PMID: 22500836 Review.
Cited by
-
Pre-encoded responsiveness to type I interferon in the peripheral immune system defines outcome of PD1 blockade therapy.Nat Immunol. 2022 Aug;23(8):1273-1283. doi: 10.1038/s41590-022-01262-7. Epub 2022 Jul 14. Nat Immunol. 2022. PMID: 35835962
-
The regulation of Tfh cell differentiation by β-hydroxybutyrylation modification of transcription factor Bcl6.Chromosoma. 2023 Nov;132(4):257-268. doi: 10.1007/s00412-023-00799-2. Epub 2023 May 25. Chromosoma. 2023. PMID: 37227491 Free PMC article.
-
Harnessing T Follicular Helper Cell Responses for HIV Vaccine Development.Viruses. 2018 Jun 19;10(6):336. doi: 10.3390/v10060336. Viruses. 2018. PMID: 29921828 Free PMC article. Review.
-
Transcription tipping points for T follicular helper cell and T-helper 1 cell fate commitment.Cell Mol Immunol. 2021 Mar;18(3):528-538. doi: 10.1038/s41423-020-00554-y. Epub 2020 Sep 30. Cell Mol Immunol. 2021. PMID: 32999454 Free PMC article. Review.
-
BCL6 represses antiviral resistance in follicular T helper cells.J Leukoc Biol. 2017 Aug;102(2):527-536. doi: 10.1189/jlb.4A1216-513RR. Epub 2017 May 26. J Leukoc Biol. 2017. PMID: 28550121 Free PMC article.
References
-
- Yu D, et al. The transcriptional repressor Bcl-6 directs T follicular helper cell lineage commitment. Immunity. 2009;31(3):457–468. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

