The genomic landscape of ribosomal peptides containing thiazole and oxazole heterocycles
- PMID: 26462797
- PMCID: PMC4603692
- DOI: 10.1186/s12864-015-2008-0
The genomic landscape of ribosomal peptides containing thiazole and oxazole heterocycles
Abstract
Background: Ribosomally synthesized and post-translationally modified peptides (RiPPs) are a burgeoning class of natural products with diverse activity that share a similar origin and common features in their biosynthetic pathways. The precursor peptides of these natural products are ribosomally produced, upon which a combination of modification enzymes installs diverse functional groups. This genetically encoded peptide-based strategy allows for rapid diversification of these natural products by mutation in the precursor genes merged with unique combinations of modification enzymes. Thiazole/oxazole-modified microcins (TOMMs) are a class of RiPPs defined by the presence of heterocycles derived from cysteine, serine, and threonine residues in the precursor peptide. TOMMs encompass a number of different families, including but not limited to the linear azol(in)e-containing peptides (streptolysin S, microcin B17, and plantazolicin), cyanobactins, thiopeptides, and bottromycins. Although many TOMMs have been explored, the increased availability of genome sequences has illuminated several unexplored TOMM producers.
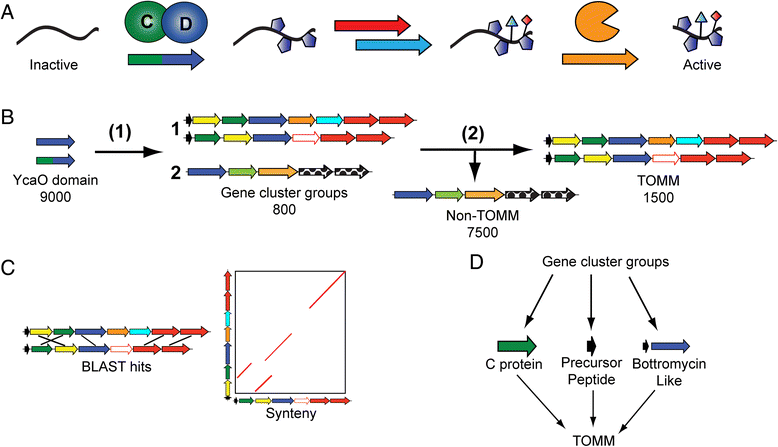
Methods: All YcaO domain-containing proteins (D protein) and the surrounding genomic regions were were obtained from the European Molecular Biology Laboratory (EMBL) and the European Bioinformatics Institute (EBI). MultiGeneBlast was used to group gene clusters contain a D protein. A number of techniques were used to identify TOMM biosynthetic gene clusters from the D protein containing gene clusters. Precursor peptides from these gene clusters were also identified. Both sequence similarity and phylogenetic analysis were used to classify the 20 diverse TOMM clusters identified.
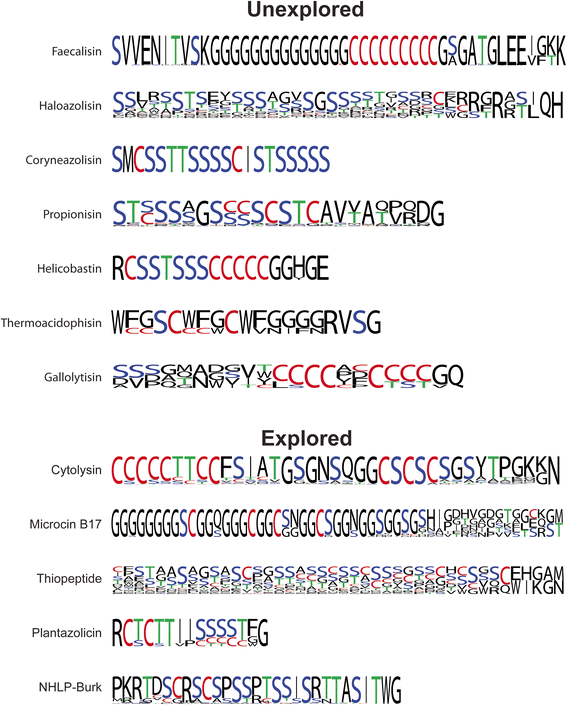
Results: Given the remarkable structural and functional diversity displayed by known TOMMs, a comprehensive bioinformatic study to catalog and classify the entire RiPP class was undertaken. Here we report the bioinformatic characterization of nearly 1,500 TOMM gene clusters from genomes in the European Molecular Biology Laboratory (EMBL) and the European Bioinformatics Institute (EBI) sequence repository. Genome mining suggests a complex diversification of modification enzymes and precursor peptides to create more than 20 distinct families of TOMMs, nine of which have not heretofore been described. Many of the identified TOMM families have an abundance of diverse precursor peptide sequences as well as unfamiliar combinations of modification enzymes, signifying a potential wealth of novel natural products on known and unknown biosynthetic scaffolds. Phylogenetic analysis suggests a widespread distribution of TOMMs across multiple phyla; however, producers of similar TOMMs are generally found in the same phylum with few exceptions.
Conclusions: The comprehensive genome mining study described herein has uncovered a myriad of unique TOMM biosynthetic clusters and provides an atlas to guide future discovery efforts. These biosynthetic gene clusters are predicted to produce diverse final products, and the identification of additional combinations of modification enzymes could expand the potential of combinatorial natural product biosynthesis.
Figures




References
-
- Arnison PG, Bibb MJ, Bierbaum G, Bowers AA, Bugni TS, Bulaj G, Camarero JA, Campopiano DJ, Challis GL, Clardy J, Cotter PD, Craik DJ, Dawson M, Dittmann E, Donadio S, Dorrestein PC, Entian KD, Fischbach MA, Garavelli JS, Goransson U, Gruber CW, Haft DH, Hemscheidt TK, Hertweck C, Hill C, Horswill AR, Jaspars M, Kelly WL, Klinman JP, Kuipers OP, et al. Ribosomally synthesized and post-translationally modified peptide natural products: overview and recommendations for a universal nomenclature. Nat Prod Rep. 2013;30(1):108–60. doi: 10.1039/C2NP20085F. - DOI - PMC - PubMed
-
- Li B, Sher D, Kelly L, Shi Y, Huang K, Knerr PJ, Joewono I, Rusch D, Chisholm SW, van der Donk WA. Catalytic promiscuity in the biosynthesis of cyclic peptide secondary metabolites in planktonic marine cyanobacteria. Proc Natl Acad Sci U S A. 2010;107(23):10430–5. doi: 10.1073/pnas.0913677107. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

