Defining the robust behaviour of the plant clock gene circuit with absolute RNA timeseries and open infrastructure
- PMID: 26468131
- PMCID: PMC4632509
- DOI: 10.1098/rsob.150042
Defining the robust behaviour of the plant clock gene circuit with absolute RNA timeseries and open infrastructure
Abstract
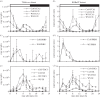
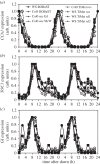
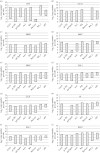
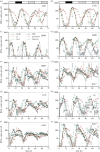
Our understanding of the complex, transcriptional feedback loops in the circadian clock mechanism has depended upon quantitative, timeseries data from disparate sources. We measure clock gene RNA profiles in Arabidopsis thaliana seedlings, grown with or without exogenous sucrose, or in soil-grown plants and in wild-type and mutant backgrounds. The RNA profiles were strikingly robust across the experimental conditions, so current mathematical models are likely to be broadly applicable in leaf tissue. In addition to providing reference data, unexpected behaviours included co-expression of PRR9 and ELF4, and regulation of PRR5 by GI. Absolute RNA quantification revealed low levels of PRR9 transcripts (peak approx. 50 copies cell(-1)) compared with other clock genes, and threefold higher levels of LHY RNA (more than 1500 copies cell(-1)) than of its close relative CCA1. The data are disseminated from BioDare, an online repository for focused timeseries data, which is expected to benefit mechanistic modelling. One data subset successfully constrained clock gene expression in a complex model, using publicly available software on parallel computers, without expert tuning or programming. We outline the empirical and mathematical justification for data aggregation in understanding highly interconnected, dynamic networks such as the clock, and the observed design constraints on the resources required to make this approach widely accessible.
Keywords: biological clocks; circadian rhythms; data management; gene regulatory networks; model optimization; plant biology.
© 2015 The Authors.
Figures


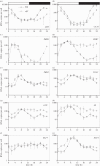



References
-
- Zhang EE, Kay SA. 2010. Clocks not winding down: unravelling circadian networks. Nat. Rev. Mol. Cell Biol. 11, 764–776. (doi:10.1038/nrm2995) - DOI - PubMed
-
- Dong G, Golden SS. 2008. How a cyanobacterium tells time. Curr. Opin. Microbiol. 11, 541–546. (doi:10.1016/j.mib.2008.10.003) - DOI - PMC - PubMed
-
- Dodd AN, et al. 2008. Plant circadian clocks increase photosynthesis, growth, survival, and competitive advantage. Science 309, 630–633. (doi:10.1126/science.1115581) - DOI - PubMed
-
- Ouyang Y, Andersson CR, Kondo T, Golden SS, Johnson CH. 1998. Resonating circadian clocks enhance fitness in cyanobacteria. Proc. Natl Acad. Sci. USA 95, 8660–8664. (doi:10.1073/pnas.95.15.8660) - DOI - PMC - PubMed
-
- Kinmonth-Schultz HA, Golembeski GS, Imaizumi T. 2013. Circadian clock-regulated physiological outputs: dynamic responses in nature. Semin. Cell Dev. Biol. 24, 407–413. (doi:10.1016/j.semcdb.2013.02.006) - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
