Detection of Diffusion Heterogeneity in Single Particle Tracking Trajectories Using a Hidden Markov Model with Measurement Noise Propagation
- PMID: 26473352
- PMCID: PMC4608688
- DOI: 10.1371/journal.pone.0140759
Detection of Diffusion Heterogeneity in Single Particle Tracking Trajectories Using a Hidden Markov Model with Measurement Noise Propagation
Abstract
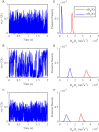
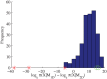
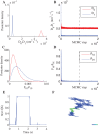
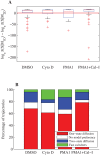
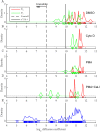
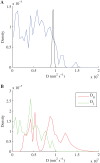
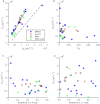
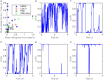
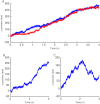
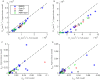
We develop a Bayesian analysis framework to detect heterogeneity in the diffusive behaviour of single particle trajectories on cells, implementing model selection to classify trajectories as either consistent with Brownian motion or with a two-state (diffusion coefficient) switching model. The incorporation of localisation accuracy is essential, as otherwise false detection of switching within a trajectory was observed and diffusion coefficient estimates were inflated. Since our analysis is on a single trajectory basis, we are able to examine heterogeneity between trajectories in a quantitative manner. Applying our method to the lymphocyte function-associated antigen 1 (LFA-1) receptor tagged with latex beads (4 s trajectories at 1000 frames s(-1)), both intra- and inter-trajectory heterogeneity were detected; 12-26% of trajectories display clear switching between diffusive states dependent on condition, whilst the inter-trajectory variability is highly structured with the diffusion coefficients being related by D1 = 0.68D0 - 1.5 × 10(4) nm2 s(-1), suggestive that on these time scales we are detecting switching due to a single process. Further, the inter-trajectory variability of the diffusion coefficient estimates (1.6 × 10(2) - 2.6 × 10(5) nm2 s(-1)) is very much larger than the measurement uncertainty within trajectories, suggesting that LFA-1 aggregation and cytoskeletal interactions are significantly affecting mobility, whilst the timescales of these processes are distinctly different giving rise to inter- and intra-trajectory variability. There is also an 'immobile' state (defined as D < 3.0 × 103 nm2 s-1) that is rarely involved in switching, immobility occurring with the highest frequency (47%) under T cell activation (phorbol-12-myristate-13-acetate (PMA) treatment) with enhanced cytoskeletal attachment (calpain inhibition). Such 'immobile' states frequently display slow linear drift, potentially reflecting binding to a dynamic actin cortex. Our methods allow significantly more information to be extracted from individual trajectories (ultimately limited by time resolution and time-series length), and allow statistical comparisons between trajectories thereby quantifying inter-trajectory heterogeneity. Such methods will be highly informative for the construction and fitting of molecule mobility models within membranes incorporating aggregation, binding to the cytoskeleton, or traversing membrane microdomains.
Conflict of interest statement
Figures


Similar articles
-
Trajectory Analysis in Single-Particle Tracking: From Mean Squared Displacement to Machine Learning Approaches.Int J Mol Sci. 2024 Aug 8;25(16):8660. doi: 10.3390/ijms25168660. Int J Mol Sci. 2024. PMID: 39201346 Free PMC article. Review.
-
A hidden Markov model for single particle tracks quantifies dynamic interactions between LFA-1 and the actin cytoskeleton.PLoS Comput Biol. 2009 Nov;5(11):e1000556. doi: 10.1371/journal.pcbi.1000556. Epub 2009 Nov 6. PLoS Comput Biol. 2009. PMID: 19893741 Free PMC article.
-
Cytoskeletal regulation couples LFA-1 conformational changes to receptor lateral mobility and clustering.Immunity. 2006 Aug;25(2):297-308. doi: 10.1016/j.immuni.2006.06.012. Epub 2006 Aug 10. Immunity. 2006. PMID: 16901728
-
A Hidden Markov Model for Detecting Confinement in Single-Particle Tracking Trajectories.Biophys J. 2018 Nov 6;115(9):1741-1754. doi: 10.1016/j.bpj.2018.09.005. Epub 2018 Sep 13. Biophys J. 2018. PMID: 30274829 Free PMC article.
-
Regulation of T-cell activation by the cytoskeleton.Nat Rev Immunol. 2007 Feb;7(2):131-43. doi: 10.1038/nri2021. Nat Rev Immunol. 2007. PMID: 17259969 Review.
Cited by
-
Trajectory Analysis in Single-Particle Tracking: From Mean Squared Displacement to Machine Learning Approaches.Int J Mol Sci. 2024 Aug 8;25(16):8660. doi: 10.3390/ijms25168660. Int J Mol Sci. 2024. PMID: 39201346 Free PMC article. Review.
-
Pointwise error estimates in localization microscopy.Nat Commun. 2017 May 3;8:15115. doi: 10.1038/ncomms15115. Nat Commun. 2017. PMID: 28466844 Free PMC article.
-
Bayesian Inference: The Comprehensive Approach to Analyzing Single-Molecule Experiments.Annu Rev Biophys. 2021 May 6;50:191-208. doi: 10.1146/annurev-biophys-082120-103921. Epub 2021 Feb 3. Annu Rev Biophys. 2021. PMID: 33534607 Free PMC article. Review.
-
Ligand Separation on Nanoconstructs Affects Targeting Selectivity to Protein Dimers on Cell Membranes.Nano Lett. 2024 Jan 10;24(1):519-524. doi: 10.1021/acs.nanolett.3c04641. Epub 2023 Dec 21. Nano Lett. 2024. PMID: 38126338 Free PMC article.
-
Impact of Feature Choice on Machine Learning Classification of Fractional Anomalous Diffusion.Entropy (Basel). 2020 Dec 19;22(12):1436. doi: 10.3390/e22121436. Entropy (Basel). 2020. PMID: 33352694 Free PMC article.
References
-
- Lingwood D, Simons K. Lipid rafts as a membrane-organizing principle. Science. 2010. January;327(5961):46–50. - PubMed
-
- Kusumi A, Nakada C, Ritchie K, Murase K, Suzuki K, Murakoshi H, et al. Paradigm shift of the plasma membrane concept from the two-dimensional continuum fluid to the partitioned fluid: high-speed single-molecule tracking of membrane molecules. Annual review of biophysics and biomolecular structure. 2005;34:351–378. 10.1146/annurev.biophys.34.040204.144637 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

