AB-Bind: Antibody binding mutational database for computational affinity predictions
- PMID: 26473627
- PMCID: PMC4815335
- DOI: 10.1002/pro.2829
AB-Bind: Antibody binding mutational database for computational affinity predictions
Abstract
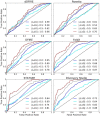
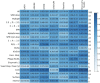
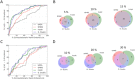
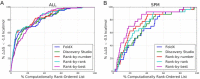
Antibodies (Abs) are a crucial component of the immune system and are often used as diagnostic and therapeutic agents. The need for high-affinity and high-specificity antibodies in research and medicine is driving the development of computational tools for accelerating antibody design and discovery. We report a diverse set of antibody binding data with accompanying structures that can be used to evaluate methods for modeling antibody interactions. Our Antibody-Bind (AB-Bind) database includes 1101 mutants with experimentally determined changes in binding free energies (ΔΔG) across 32 complexes. Using the AB-Bind data set, we evaluated the performance of protein scoring potentials in their ability to predict changes in binding free energies upon mutagenesis. Numerical correlations between computed and observed ΔΔG values were low (r = 0.16-0.45), but the potentials exhibited predictive power for classifying variants as improved vs weakened binders. Performance was evaluated using the area under the curve (AUC) for receiver operator characteristic (ROC) curves; the highest AUC values for 527 mutants with |ΔΔG| > 1.0 kcal/mol were 0.81, 0.87, and 0.88 using STATIUM, FoldX, and Discovery Studio scoring potentials, respectively. Some methods could also enrich for variants with improved binding affinity; FoldX and Discovery Studio were able to correctly rank 42% and 30%, respectively, of the 80 most improved binders (those with ΔΔG < -1.0 kcal/mol) in the top 5% of the database. This modest predictive performance has value but demonstrates the continuing need to develop and improve protein energy functions for affinity prediction.
Keywords: affinity optimization; antibody affinity; antibody mutagenesis; computational affinity prediction; mutational database; protein interface design; protein-protein interactions; scoring interface mutations; structure-based modeling.
© 2015 The Protein Society.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

