Targeting autocrine HB-EGF signaling with specific ADAM12 inhibition using recombinant ADAM12 prodomain
- PMID: 26477568
- PMCID: PMC4609913
- DOI: 10.1038/srep15150
Targeting autocrine HB-EGF signaling with specific ADAM12 inhibition using recombinant ADAM12 prodomain
Abstract
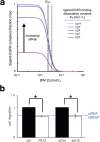
Dysregulation of ErbB-family signaling underlies numerous pathologies and has been therapeutically targeted through inhibiting ErbB-receptors themselves or their cognate ligands. For the latter, "decoy" antibodies have been developed to sequester ligands including heparin-binding epidermal growth factor (HB-EGF); however, demonstrating sufficient efficacy has been difficult. Here, we hypothesized that this strategy depends on properties such as ligand-receptor binding affinity, which varies widely across the known ErbB-family ligands. Guided by computational modeling, we found that high-affinity ligands such as HB-EGF are more difficult to target with decoy antibodies compared to low-affinity ligands such as amphiregulin (AREG). To address this issue, we developed an alternative method for inhibiting HB-EGF activity by targeting its cleavage from the cell surface. In a model of the invasive disease endometriosis, we identified A Disintegrin and Metalloproteinase 12 (ADAM12) as a protease implicated in HB-EGF shedding. We designed a specific inhibitor of ADAM12 based on its recombinant prodomain (PA12), which selectively inhibits ADAM12 but not ADAM10 or ADAM17. In endometriotic cells, PA12 significantly reduced HB-EGF shedding and resultant cellular migration. Overall, specific inhibition of ligand shedding represents a possible alternative to decoy antibodies, especially for ligands such as HB-EGF that exhibit high binding affinity and localized signaling.
Figures




Similar articles
-
Mitogen-activated protein kinase (MAPK) regulates leukotriene D4-induced HB-EGF and ADAM12 expression in human airway smooth muscle cells.Asian Pac J Allergy Immunol. 2013 Mar;31(1):58-66. Asian Pac J Allergy Immunol. 2013. PMID: 23517395
-
[Membrane-anchored heparin-binding EGF-like growth factor processing by ADAM12 in cardiac hypertrophy].Nihon Rinsho. 2003 May;61(5):767-75. Nihon Rinsho. 2003. PMID: 12755001 Review. Japanese.
-
Cardiac hypertrophy is inhibited by antagonism of ADAM12 processing of HB-EGF: metalloproteinase inhibitors as a new therapy.Nat Med. 2002 Jan;8(1):35-40. doi: 10.1038/nm0102-35. Nat Med. 2002. PMID: 11786904
-
EGF promotes the shedding of soluble E-cadherin in an ADAM10-dependent manner in prostate epithelial cells.Cell Signal. 2012 Feb;24(2):532-538. doi: 10.1016/j.cellsig.2011.10.004. Epub 2011 Oct 14. Cell Signal. 2012. PMID: 22024284 Free PMC article.
-
ADAM-mediated ectodomain shedding of HB-EGF in receptor cross-talk.Biochim Biophys Acta. 2005 Aug 1;1751(1):110-7. doi: 10.1016/j.bbapap.2004.11.009. Epub 2004 Dec 8. Biochim Biophys Acta. 2005. PMID: 16054021 Review.
Cited by
-
Imaging the pharmacology of nanomaterials by intravital microscopy: Toward understanding their biological behavior.Adv Drug Deliv Rev. 2017 Apr;113:61-86. doi: 10.1016/j.addr.2016.05.023. Epub 2016 Jun 4. Adv Drug Deliv Rev. 2017. PMID: 27266447 Free PMC article. Review.
-
Single-Cell Intravital Microscopy of Trastuzumab Quantifies Heterogeneous in vivo Kinetics.Cytometry A. 2020 May;97(5):528-539. doi: 10.1002/cyto.a.23872. Epub 2019 Aug 19. Cytometry A. 2020. PMID: 31423731 Free PMC article.
-
MenaINV mediates synergistic cross-talk between signaling pathways driving chemotaxis and haptotaxis.Mol Biol Cell. 2016 Oct 15;27(20):3085-3094. doi: 10.1091/mbc.E16-04-0212. Epub 2016 Aug 24. Mol Biol Cell. 2016. PMID: 27559126 Free PMC article.
-
Regulation of Fibrotic Processes in the Liver by ADAM Proteases.Cells. 2019 Oct 9;8(10):1226. doi: 10.3390/cells8101226. Cells. 2019. PMID: 31601007 Free PMC article. Review.
-
Identification of ADAM12 as a Novel Basigin Sheddase.Int J Mol Sci. 2019 Apr 22;20(8):1957. doi: 10.3390/ijms20081957. Int J Mol Sci. 2019. PMID: 31013576 Free PMC article.
References
-
- Wilson T. R., Lee D. Y., Berry L., Shames D. S. & Settleman J. Neuregulin-1-mediated autocrine signaling underlies sensitivity to HER2 kinase inhibitors in a subset of human cancers. Cancer Cell 20, 158–172 (2011). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

