Crawling and Gliding: A Computational Model for Shape-Driven Cell Migration
- PMID: 26488304
- PMCID: PMC4619082
- DOI: 10.1371/journal.pcbi.1004280
Crawling and Gliding: A Computational Model for Shape-Driven Cell Migration
Abstract
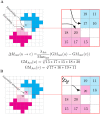
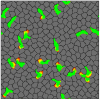
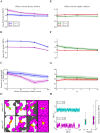
Cell migration is a complex process involving many intracellular and extracellular factors, with different cell types adopting sometimes strikingly different morphologies. Modeling realistically behaving cells in tissues is computationally challenging because it implies dealing with multiple levels of complexity. We extend the Cellular Potts Model with an actin-inspired feedback mechanism that allows small stochastic cell rufflings to expand to cell protrusions. This simple phenomenological model produces realistically crawling and deforming amoeboid cells, and gliding half-moon shaped keratocyte-like cells. Both cell types can migrate randomly or follow directional cues. They can squeeze in between other cells in densely populated environments or migrate collectively. The model is computationally light, which allows the study of large, dense and heterogeneous tissues containing cells with realistic shapes and migratory properties.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures









Similar articles
-
Viscoelastic gel-strip model for the simulation of migrating cells.Ann Biomed Eng. 2011 Nov;39(11):2735-49. doi: 10.1007/s10439-011-0360-z. Epub 2011 Jul 29. Ann Biomed Eng. 2011. PMID: 21800204
-
Cortical factor feedback model for cellular locomotion and cytofission.PLoS Comput Biol. 2009 Mar;5(3):e1000310. doi: 10.1371/journal.pcbi.1000310. Epub 2009 Mar 13. PLoS Comput Biol. 2009. PMID: 19282961 Free PMC article.
-
Linking morphodynamics and directional persistence of T lymphocyte migration.J R Soc Interface. 2015 May 6;12(106):20141412. doi: 10.1098/rsif.2014.1412. J R Soc Interface. 2015. PMID: 25904526 Free PMC article.
-
Crawling toward a unified model of cell mobility: spatial and temporal regulation of actin dynamics.Annu Rev Biochem. 2004;73:209-39. doi: 10.1146/annurev.biochem.73.011303.073844. Annu Rev Biochem. 2004. PMID: 15189141 Review.
-
Amoeboid leukocyte crawling through extracellular matrix: lessons from the Dictyostelium paradigm of cell movement.J Leukoc Biol. 2001 Oct;70(4):491-509. J Leukoc Biol. 2001. PMID: 11590185 Review.
Cited by
-
Interpreting T-cell search "strategies" in the light of evolution under constraints.PLoS Comput Biol. 2023 Feb 27;19(2):e1010918. doi: 10.1371/journal.pcbi.1010918. eCollection 2023 Feb. PLoS Comput Biol. 2023. PMID: 36848395 Free PMC article.
-
Collective Cell Radial Ordered Migration in Spatial Confinement.Adv Sci (Weinh). 2024 May;11(20):e2307487. doi: 10.1002/advs.202307487. Epub 2024 Mar 23. Adv Sci (Weinh). 2024. PMID: 38520715 Free PMC article.
-
Density-Dependent Migration Characteristics of Cancer Cells Driven by Pseudopod Interaction.Front Cell Dev Biol. 2022 Apr 25;10:854721. doi: 10.3389/fcell.2022.854721. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 35547818 Free PMC article.
-
A minimal computational model for three-dimensional cell migration.J R Soc Interface. 2019 Dec;16(161):20190619. doi: 10.1098/rsif.2019.0619. Epub 2019 Dec 18. J R Soc Interface. 2019. PMID: 31847757 Free PMC article.
-
Hybrid cellular Potts and bead-spring modeling of cells in fibrous extracellular matrix.Biophys J. 2023 Jul 11;122(13):2609-2622. doi: 10.1016/j.bpj.2023.05.013. Epub 2023 May 13. Biophys J. 2023. PMID: 37183398 Free PMC article.
References
-
- Friedl P, Wolf K. Plasticity of cell migration: a multiscale tuning model. The Journal of Cell Biology. 2010;188(1):11–19. Available from: http://jcb.rupress.org/content/188/1/11.abstract. 10.1083/jcb.200909003 - DOI - PMC - PubMed
-
- Lämmermann T, Sixt M. Mechanical modes of ‘amoeboid’ cell migration. Current Opinion in Cell Biology. 2009;21(5):636–644. Available from: http://www.sciencedirect.com/science/article/pii/S0955067409001148. 10.1016/j.ceb.2009.05.003 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

