RNA Silencing May Play a Role in but Is Not the Only Determinant of the Multiplicity of Infection
- PMID: 26491166
- PMCID: PMC4702557
- DOI: 10.1128/JVI.02345-15
RNA Silencing May Play a Role in but Is Not the Only Determinant of the Multiplicity of Infection
Abstract
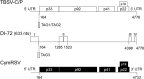
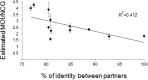
The multiplicity of infection (MOI), i.e., the number of viral genomes that infect a cell, is an important parameter in virus evolution, which for each virus and environment may have an optimum value that maximizes virus fitness. Thus, the MOI might be controlled by virus functions, an underexplored hypothesis in eukaryote-infecting viruses. To analyze if the MOI is controlled by virus functions, we estimated the MOI in plants coinfected by two genetic variants of Tomato bushy stunt virus (TBSV); by TBSV and a TBSV-derived defective interfering RNA (DI-RNA); or by TBSV and a second tombusvirus, Cymbidium ringspot virus (CymRSV). The MOI was significantly larger in TBSV-CymRSV coinfections (~4.0) than in TBSV-TBSV or TBSV-DI-RNA coinfections (~1.7 to 2.2). Coinfections by CymRSV or TBSV with chimeras in which an open reading frame (ORF) of one virus species was replaced by that of the other identified a role of viral proteins in determining the MOI, which ranged from 1.6 to 3.9 depending on the coinfecting genotypes. However, no virus-encoded protein or genomic region was the sole MOI determinant. Coinfections by CymRSV and TBSV mutants in which the expression of the gene-silencing suppressor protein p19 was abolished also showed a possible role of gene silencing in MOI determination. Taken together, these results demonstrate that the MOI is a quantitative trait showing continuous variation and that as such it has a complex determination involving different virus-encoded functions.
Importance: The number of viral genomes infecting a cell, or the multiplicity of infection (MOI), is an important parameter in virus evolution affecting recombination rates, selection intensity on viral genes, evolution of multipartite genomes, or hyperparasitism by satellites or defective interfering particles. For each virus and environment, the MOI may have an optimum value that maximizes virus fitness, but little is known about MOI control in eukaryote-infecting viruses. We show here that in plants coinfected by two genotypes of Tomato bushy stunt virus (TBSV), the MOI was lower than in plants coinfected by TBSV and Cymbidium ringspot virus (CymRSV). Coinfections by CymRSV or TBSV with TBSV-CymRSV chimeras showed a role of viral proteins in MOI determination. Coinfections by CymRSV and TBSV mutants not expressing the gene-silencing suppressor protein also showed a role of gene silencing in MOI determination. The results demonstrate that the MOI is a quantitative trait with a complex determination involving different viral functions.
Copyright © 2015, American Society for Microbiology. All Rights Reserved.
Figures

Similar articles
-
Properties of a novel satellite RNA associated with tomato bushy stunt virus infections.J Gen Virol. 2010 Sep;91(Pt 9):2393-401. doi: 10.1099/vir.0.022046-0. Epub 2010 May 19. J Gen Virol. 2010. PMID: 20484559
-
A novel co-delivery system consisting of a Tomato bushy stunt virus and a defective interfering RNA for studying gene silencing.J Virol Methods. 2003 Jul;111(1):37-42. doi: 10.1016/s0166-0934(03)00149-6. J Virol Methods. 2003. PMID: 12821195
-
A survey of resistance to Tomato bushy stunt virus in the genus Nicotiana reveals that the hypersensitive response is triggered by one of three different viral proteins.Mol Plant Microbe Interact. 2013 Feb;26(2):240-8. doi: 10.1094/MPMI-06-12-0157-R. Mol Plant Microbe Interact. 2013. PMID: 23075040
-
Global genomics and proteomics approaches to identify host factors as targets to induce resistance against Tomato bushy stunt virus.Adv Virus Res. 2010;76:123-77. doi: 10.1016/S0065-3527(10)76004-8. Epub 2010 Mar 31. Adv Virus Res. 2010. PMID: 20965073 Free PMC article. Review.
-
The roles of host factors in tombusvirus RNA recombination.Adv Virus Res. 2011;81:63-84. doi: 10.1016/B978-0-12-385885-6.00008-0. Adv Virus Res. 2011. PMID: 22094079 Review.
Cited by
-
Bottleneck, Isolate, Amplify, Select (BIAS) as a mechanistic framework for intracellular population dynamics of positive-sense RNA viruses.Virus Evol. 2020 Nov 12;6(2):veaa086. doi: 10.1093/ve/veaa086. eCollection 2020 Jul. Virus Evol. 2020. PMID: 33343926 Free PMC article. Review.
-
The role of co-opted ESCRT proteins and lipid factors in protection of tombusviral double-stranded RNA replication intermediate against reconstituted RNAi in yeast.PLoS Pathog. 2017 Jul 31;13(7):e1006520. doi: 10.1371/journal.ppat.1006520. eCollection 2017 Jul. PLoS Pathog. 2017. PMID: 28759634 Free PMC article.
References
-
- Escriu F, Fraile A, García-Arenal F. 2003. The evolution of virulence in a plant virus. Evolution 57:755–765. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

