Evidence of Selection against Complex Mitotic-Origin Aneuploidy during Preimplantation Development
- PMID: 26491874
- PMCID: PMC4619652
- DOI: 10.1371/journal.pgen.1005601
Evidence of Selection against Complex Mitotic-Origin Aneuploidy during Preimplantation Development
Abstract
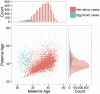
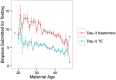
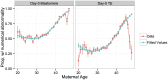
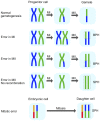
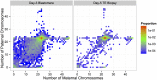
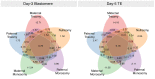
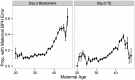
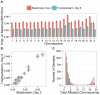
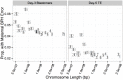
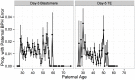
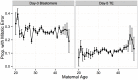
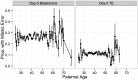
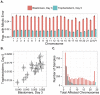
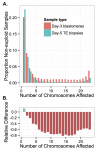
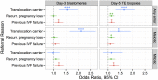
Whole-chromosome imbalances affect over half of early human embryos and are the leading cause of pregnancy loss. While these errors frequently arise in oocyte meiosis, many such whole-chromosome abnormalities affecting cleavage-stage embryos are the result of chromosome missegregation occurring during the initial mitotic cell divisions. The first wave of zygotic genome activation at the 4-8 cell stage results in the arrest of a large proportion of embryos, the vast majority of which contain whole-chromosome abnormalities. Thus, the full spectrum of meiotic and mitotic errors can only be detected by sampling after the initial cell divisions, but prior to this selective filter. Here, we apply 24-chromosome preimplantation genetic screening (PGS) to 28,052 single-cell day-3 blastomere biopsies and 18,387 multi-cell day-5 trophectoderm biopsies from 6,366 in vitro fertilization (IVF) cycles. We precisely characterize the rates and patterns of whole-chromosome abnormalities at each developmental stage and distinguish errors of meiotic and mitotic origin without embryo disaggregation, based on informative chromosomal signatures. We show that mitotic errors frequently involve multiple chromosome losses that are not biased toward maternal or paternal homologs. This outcome is characteristic of spindle abnormalities and chaotic cell division detected in previous studies. In contrast to meiotic errors, our data also show that mitotic errors are not significantly associated with maternal age. PGS patients referred due to previous IVF failure had elevated rates of mitotic error, while patients referred due to recurrent pregnancy loss had elevated rates of meiotic error, controlling for maternal age. These results support the conclusion that mitotic error is the predominant mechanism contributing to pregnancy losses occurring prior to blastocyst formation. This high-resolution view of the full spectrum of whole-chromosome abnormalities affecting early embryos provides insight into the cytogenetic mechanisms underlying their formation and the consequences for human fertility.
Conflict of interest statement
I have read the journal’s policy and the authors of this manuscript have the following competing interests: DAP has received stock options in Natera, Inc., as consulting fees. ZPD, AR, MB, MH, SS, and MR are full-time employees of and hold stock or options to hold stock in Natera, Inc. Stanford University filed a provisional patent related to this work with the U.S. Patent and Trademark Office on 14 November 2014 (USSN 62/080,251).
Figures

Similar articles
-
Sequential comprehensive chromosome analysis on polar bodies, blastomeres and trophoblast: insights into female meiotic errors and chromosomal segregation in the preimplantation window of embryo development.Hum Reprod. 2013 Feb;28(2):509-18. doi: 10.1093/humrep/des394. Epub 2012 Nov 11. Hum Reprod. 2013. PMID: 23148203
-
The incidence and origin of segmental aneuploidy in human oocytes and preimplantation embryos.Hum Reprod. 2017 Dec 1;32(12):2549-2560. doi: 10.1093/humrep/dex324. Hum Reprod. 2017. PMID: 29126206
-
Origins and rates of aneuploidy in human blastomeres.Fertil Steril. 2012 Feb;97(2):395-401. doi: 10.1016/j.fertnstert.2011.11.034. Epub 2011 Dec 21. Fertil Steril. 2012. PMID: 22195772
-
The cytogenetic constitution of human blastocysts: insights from comprehensive chromosome screening strategies.Hum Reprod Update. 2019 Jan 1;25(1):15-33. doi: 10.1093/humupd/dmy036. Hum Reprod Update. 2019. PMID: 30395265 Review.
-
The genetics of preimplantation embryonic arrest: the role of aneuploidies.Curr Opin Obstet Gynecol. 2025 Jun 1;37(3):123-129. doi: 10.1097/GCO.0000000000001020. Epub 2025 Apr 28. Curr Opin Obstet Gynecol. 2025. PMID: 40167998 Review.
Cited by
-
Consequences of Chromosome Loss: Why Do Cells Need Each Chromosome Twice?Cells. 2022 May 3;11(9):1530. doi: 10.3390/cells11091530. Cells. 2022. PMID: 35563836 Free PMC article. Review.
-
The importance of standardizing criteria for PGT-A interpretation of blastocysts analyzed by next-generation sequencing.JBRA Assist Reprod. 2023 Sep 12;27(3):453-462. doi: 10.5935/1518-0557.20230011. JBRA Assist Reprod. 2023. PMID: 37579264 Free PMC article.
-
Female-age-dependent changes in the lipid fingerprint of the mammalian oocytes.Hum Reprod. 2024 Dec 1;39(12):2754-2767. doi: 10.1093/humrep/deae225. Hum Reprod. 2024. PMID: 39366679 Free PMC article.
-
Maternal selection of human embryos in early gestation: Insights from recurrent miscarriage.Semin Cell Dev Biol. 2022 Nov;131:14-24. doi: 10.1016/j.semcdb.2022.01.007. Epub 2022 Jan 31. Semin Cell Dev Biol. 2022. PMID: 35094946 Free PMC article. Review.
-
Preimplantation chromosomal mosaics, chimaeras and confined placental mosaicism.Reprod Fertil. 2022 Apr 5;3(2):R66-R90. doi: 10.1530/RAF-21-0095. eCollection 2022 Apr 1. Reprod Fertil. 2022. PMID: 35514539 Free PMC article. Review.
References
-
- Edmonds DK, Lindsay KS, Miller JF, Williamson E, Wood PJ. Early embryonic mortality in women. Obstetrical & Gynecological Survey. 1983;38(7):433–434. - PubMed
-
- Levy B, Sigurjonsson S, Petterson B, Maisenbacher M, Hall M, Demko Z, et al. Genomic imbalance in products of conception: SNP chromosomal microarray analysis. Obstetrics & Gynecology. 2014;124(2pt1):202–209. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

