S-nitrosylation triggers ABI5 degradation to promote seed germination and seedling growth
- PMID: 26493030
- PMCID: PMC4639896
- DOI: 10.1038/ncomms9669
S-nitrosylation triggers ABI5 degradation to promote seed germination and seedling growth
Abstract
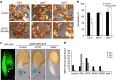
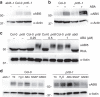
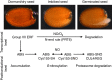
Plant survival depends on seed germination and progression through post-germinative developmental checkpoints. These processes are controlled by the stress phytohormone abscisic acid (ABA). ABA regulates the basic leucine zipper transcriptional factor ABI5, a central hub of growth repression, while the reactive nitrogen molecule nitric oxide (NO) counteracts ABA during seed germination. However, the molecular mechanisms by which seeds sense more favourable conditions and start germinating have remained elusive. Here we show that ABI5 promotes growth via NO, and that ABI5 accumulation is altered in genetic backgrounds with impaired NO homeostasis. S-nitrosylation of ABI5 at cysteine-153 facilitates its degradation through CULLIN4-based and KEEP ON GOING E3 ligases, and promotes seed germination. Conversely, mutation of ABI5 at cysteine-153 deregulates protein stability and inhibition of seed germination by NO depletion. These findings suggest an inverse molecular link between NO and ABA hormone signalling through distinct posttranslational modifications of ABI5 during early seedling development.
Figures



References
-
- Nambara E. et al.. Abscisic acid and the control of seed dormancy and germination. Seed Sci. Res. 20, 55–67 (2010).
-
- Lopez-Molina L., Mongrand S., McLachlin D., Chait B. & Chua N. H. ABI5 acts downstream of ABI3 to execute an ABA-dependent growth arrest during germination. Plant J. 32, 1–12 (2002). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

