D-Amino Acid Probes for Penicillin Binding Protein-based Bacterial Surface Labeling
- PMID: 26499795
- PMCID: PMC4683274
- DOI: 10.1074/jbc.M115.683342
D-Amino Acid Probes for Penicillin Binding Protein-based Bacterial Surface Labeling
Abstract
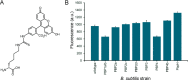
Peptidoglycan is an essential and highly conserved mesh structure that surrounds bacterial cells. It plays a critical role in retaining a defined cell shape, and, in the case of pathogenic Gram-positive bacteria, it lies at the interface between bacterial cells and the host organism. Intriguingly, bacteria can metabolically incorporate unnatural D-amino acids into the peptidoglycan stem peptide directly from the surrounding medium, a process mediated by penicillin binding proteins (PBPs). Metabolic peptidoglycan remodeling via unnatural D-amino acids has provided unique insights into peptidoglycan biosynthesis of live bacteria and has also served as the basis of a synthetic immunology strategy with potential therapeutic implications. A striking feature of this process is the vast promiscuity displayed by PBPs in tolerating entirely unnatural side chains. However, the chemical space and physical features of this side chain promiscuity have not been determined systematically. In this report, we designed and synthesized a library of variants displaying diverse side chains to comprehensively establish the tolerability of unnatural D-amino acids by PBPs in both Gram-positive and Gram-negative organisms. In addition, nine Bacillus subtilis PBP-null mutants were evaluated with the goal of identifying a potential primary PBP responsible for unnatural D-amino acid incorporation and gaining insights into the temporal control of PBP activity. We empirically established the scope of physical parameters that govern the metabolic incorporation of unnatural D-amino acids into bacterial peptidoglycan.
Keywords: Bacillus; amino acid; bacteria; bacterial conjugation; cell surface; cell wall; peptide biosynthesis; peptide chemical synthesis; peptides; peptidoglycan.
© 2015 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures








References
-
- Vollmer W., Blanot D., and de Pedro M. A. (2008) Peptidoglycan structure and architecture. FEMS Microbiol. Rev. 32, 149–167 - PubMed
-
- Frère J. M., and Joris B. (1985) Penicillin-sensitive enzymes in peptidoglycan biosynthesis. Crit. Rev. Microbiol. 11, 299–396 - PubMed
-
- van Heijenoort J. (2001) Recent advances in the formation of the bacterial peptidoglycan monomer unit. Nat. Prod. Rep. 18, 503–519 - PubMed
-
- Bugg T. D., and Walsh C. T. (1992) Intracellular steps of bacterial cell wall peptidoglycan biosynthesis: enzymology, antibiotics, and antibiotic resistance. Nat. Prod. Rep. 9, 199–215 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

