Computational reconstitution of spine calcium transients from individual proteins
- PMID: 26500546
- PMCID: PMC4595661
- DOI: 10.3389/fnsyn.2015.00017
Computational reconstitution of spine calcium transients from individual proteins
Abstract
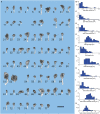
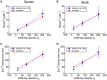
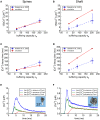
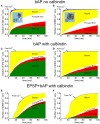
We have built a stochastic model in the program MCell that simulates Ca(2+) transients in spines from the principal molecular components believed to control Ca(2+) entry and exit. Proteins, with their kinetic models, are located within two segments of dendrites containing 88 intact spines, centered in a fully reconstructed 6 × 6 × 5 μm(3) cube of hippocampal neuropil. Protein components include AMPA- and NMDA-type glutamate receptors, L- and R-type voltage-dependent Ca(2+) channels, Na(+)/Ca(2+) exchangers, plasma membrane Ca(2+) ATPases, smooth endoplasmic reticulum Ca(2+) ATPases, immobile Ca(2+) buffers, and calbindin. Kinetic models for each protein were taken from published studies of the isolated proteins in vitro. For simulation of electrical stimuli, the time course of voltage changes in the dendritic spine was generated with the desired stimulus in the program NEURON. Voltage-dependent parameters were then continuously re-adjusted during simulations in MCell to reproduce the effects of the stimulus. Nine parameters of the model were optimized within realistic experimental limits by a process that compared results of simulations to published data. We find that simulations in the optimized model reproduce the timing and amplitude of Ca(2+) transients measured experimentally in intact neurons. Thus, we demonstrate that the characteristics of individual isolated proteins determined in vitro can accurately reproduce the dynamics of experimentally measured Ca(2+) transients in spines. The model will provide a test bed for exploring the roles of additional proteins that regulate Ca(2+) influx into spines and for studying the behavior of protein targets in the spine that are regulated by Ca(2+) influx.
Keywords: calcium channels; calcium pumps; dendritic spines; synaptic calcium; synaptic plasticity.
Figures





References
Grants and funding
- R56 MH095980/MH/NIMH NIH HHS/United States
- R01 EB000487/EB/NIBIB NIH HHS/United States
- R01 MH104319/MH/NIMH NIH HHS/United States
- R01 DA030749/DA/NIDA NIH HHS/United States
- P41 GM103712/GM/NIGMS NIH HHS/United States
- R56 EB000487/EB/NIBIB NIH HHS/United States
- R01 NS021184/NS/NINDS NIH HHS/United States
- R01 MH095980/MH/NIMH NIH HHS/United States
- R01 MH079076/MH/NIMH NIH HHS/United States
- R01 NS074644/NS/NINDS NIH HHS/United States
- P01 NS044306/NS/NINDS NIH HHS/United States
- R37 NS021184/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

