Dilp8 requires the neuronal relaxin receptor Lgr3 to couple growth to developmental timing
- PMID: 26510564
- PMCID: PMC4640092
- DOI: 10.1038/ncomms9732
Dilp8 requires the neuronal relaxin receptor Lgr3 to couple growth to developmental timing
Abstract
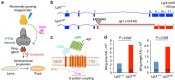
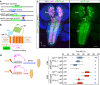
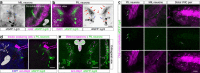
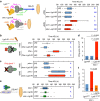
How different organs in the body sense growth perturbations in distant tissues to coordinate their size during development is poorly understood. Here we mutate an invertebrate orphan relaxin receptor gene, the Drosophila Leucine-rich repeat-containing G protein-coupled receptor 3 (Lgr3), and find body asymmetries similar to those found in insulin-like peptide 8 (dilp8) mutants, which fail to coordinate growth with developmental timing. Indeed, mutation or RNA intereference (RNAi) against Lgr3 suppresses the delay in pupariation induced by imaginal disc growth perturbation or ectopic Dilp8 expression. By tagging endogenous Lgr3 and performing cell type-specific RNAi, we map this Lgr3 activity to a new subset of CNS neurons, four of which are a pair of bilateral pars intercerebralis Lgr3-positive (PIL) neurons that respond specifically to ectopic Dilp8 by increasing cAMP-dependent signalling. Our work sheds new light on the function and evolution of relaxin receptors and reveals a novel neuroendocrine circuit responsive to growth aberrations.
Figures




References
-
- Hariharan I. K. How growth abnormalities delay ‘puberty' in Drosophila. Sci. Signal. 5, pe27 (2012). - PubMed
-
- Garelli A., Gontijo A. M., Miguela V., Caparros E. & Dominguez M. Imaginal discs secrete insulin-like peptide 8 to mediate plasticity of growth and maturation. Science 336, 579–582 (2012). - PubMed
-
- Colombani J., Andersen D. S. & Léopold P. Secreted peptide Dilp8 coordinates Drosophila tissue growth with developmental timing. Science 336, 582–585 (2012). - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

