Coevolutionary dynamics between tribe Cercopithecini tetherins and their lentiviruses
- PMID: 26531727
- PMCID: PMC4631996
- DOI: 10.1038/srep16021
Coevolutionary dynamics between tribe Cercopithecini tetherins and their lentiviruses
Abstract
Human immunodeficiency virus, a primate lentivirus (PLV), causes AIDS in humans, whereas most PLVs are less or not pathogenic in monkeys. These notions suggest that the co-evolutionary process of PLVs and their hosts associates with viral pathogenicity, and therefore, that elucidating the history of virus-host co-evolution is one of the most intriguing topics in the field of virology. To address this, recent studies have focused on the interplay between intrinsic anti-viral proteins, such as tetherin, and viral antagonists. Through an experimental-phylogenetic approach, here we investigate the co-evolutionary interplay between tribe Cercopithecini tetherin and viral antagonists, Nef and Vpu. We reveal that tribe Cercopithecini tetherins are positively selected, possibly triggered by ancient Nef-like factor(s). We reconstruct the ancestral sequence of tribe Cercopithecini tetherin and demonstrate that all Nef proteins are capable of antagonizing ancestral Cercopithecini tetherin. Further, we consider the significance of evolutionary arms race between tribe Cercopithecini and their PLVs.
Figures



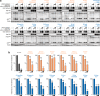

References
-
- Bailes E. et al. Hybrid origin of SIV in chimpanzees. Science 300, 1713 (2003). - PubMed
-
- Groves C. P. Order primates. Mammal species of the world: a taxonomic and geographic reference (eds Wilson D. E, Reeder D. M ), 3rd edn (Johns Hopkins University Press, 2005).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

