DDX3, a potential target for cancer treatment
- PMID: 26541825
- PMCID: PMC4636063
- DOI: 10.1186/s12943-015-0461-7
DDX3, a potential target for cancer treatment
Abstract
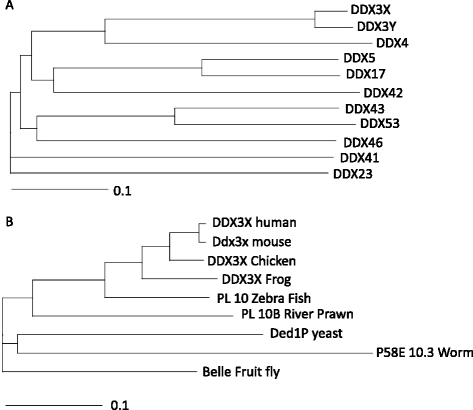
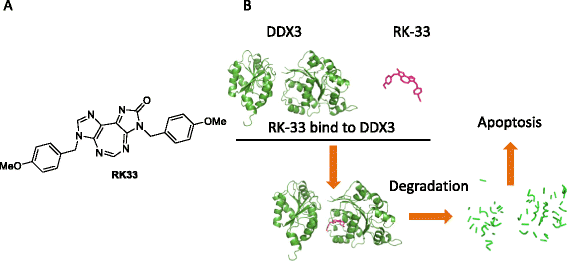
RNA helicases are a large family of proteins with a distinct motif, referred to as the DEAD/H (Asp-Glu-Ala-Asp/His). The exact functions of all the human DEAD/H box proteins are unknown. However, it has been consistently demonstrated that these proteins are associated with several aspects of energy-dependent RNA metabolism, including translation, ribosome biogenesis, and pre-mRNA splicing. In addition, DEAD/H box proteins participate in nuclear-cytoplasmic transport and organellar gene expression.A member of this RNA helicase family, DDX3, has been identified in a variety of cellular biogenesis processes, including cell-cycle regulation, cellular differentiation, cell survival, and apoptosis. In cancer, DDX3 expression has been evaluated in patient samples of breast, lung, colon, oral, and liver cancer. Both tumor suppressor and oncogenic functions have been attributed to DDX3 and are discussed in this review. In general, there is concordance with in vitro evidence to support the hypothesis that DDX3 is associated with an aggressive phenotype in human malignancies. Interestingly, very few cancer types harbor mutations in DDX3, which result in altered protein function rather than a loss of function.Efficacy of drugs to curtail cancer growth is hindered by adaptive responses that promote drug resistance, eventually leading to treatment failure. One way to circumvent development of resistant disease is to develop novel drugs that target over-expressed proteins involved in this adaptive response. Moreover, if the target gene is developmentally regulated, there is less of a possibility to abruptly accumulate mutations leading to drug resistance. In this regard, DDX3 could be a druggable target for cancer treatment. We present an overview of DDX3 biology and the currently available DDX3 inhibitors for cancer treatment.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

