Genome-wide DNA methylation profiling of CD8+ T cells shows a distinct epigenetic signature to CD4+ T cells in multiple sclerosis patients
- PMID: 26550040
- PMCID: PMC4635618
- DOI: 10.1186/s13148-015-0152-7
Genome-wide DNA methylation profiling of CD8+ T cells shows a distinct epigenetic signature to CD4+ T cells in multiple sclerosis patients
Abstract
Background: Multiple sclerosis (MS) is thought to be a T cell-mediated autoimmune disorder. MS pathogenesis is likely due to a genetic predisposition triggered by a variety of environmental factors. Epigenetics, particularly DNA methylation, provide a logical interface for environmental factors to influence the genome. In this study we aim to identify DNA methylation changes associated with MS in CD8+ T cells in 30 relapsing remitting MS patients and 28 healthy blood donors using Illumina 450K methylation arrays.
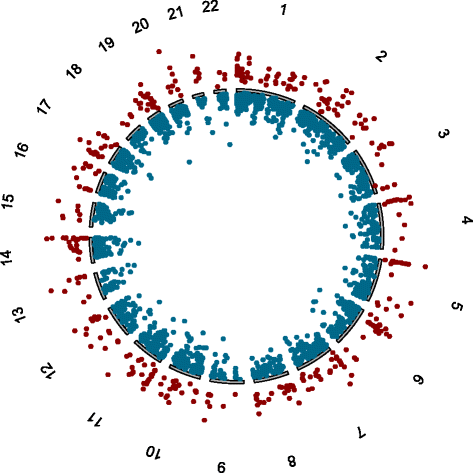
Findings: Seventy-nine differentially methylated CpGs were associated with MS. The methylation profile of CD8+ T cells was distinctive from our previously published data on CD4+ T cells in the same cohort. Most notably, there was no major CpG effect at the MS risk gene HLA-DRB1 locus in the CD8+ T cells.
Conclusion: CD8+ T cells and CD4+ T cells have distinct DNA methylation profiles. This case-control study highlights the importance of distinctive cell subtypes when investigating epigenetic changes in MS and other complex diseases.
Keywords: CD8+ T cells; DNA methylation; HLA-DRB1; Multiple sclerosis.
Figures
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

