Guidelines for the Use of Protein Domains in Acidic Phospholipid Imaging
- PMID: 26552684
- PMCID: PMC4671299
- DOI: 10.1007/978-1-4939-3170-5_15
Guidelines for the Use of Protein Domains in Acidic Phospholipid Imaging
Abstract
Acidic phospholipids are minor membrane lipids but critically important for signaling events. The main acidic phospholipids are phosphatidylinositol phosphates (PIPs also known as phosphoinositides), phosphatidylserine (PS), and phosphatidic acid (PA). Acidic phospholipids are precursors of second messengers of key signaling cascades or are second messengers themselves. They regulate the localization and activation of many proteins, and are involved in virtually all membrane trafficking events. As such, it is crucial to understand the subcellular localization and dynamics of each of these lipids within the cell. Over the years, several techniques have emerged in either fixed or live cells to analyze the subcellular localization and dynamics of acidic phospholipids. In this chapter, we review one of them: the use of genetically encoded biosensors that are based on the expression of specific lipid binding domains (LBDs) fused to fluorescent proteins. We discuss how to design such sensors, including the criteria for selecting the lipid binding domains of interest and to validate them. We also emphasize the care that must be taken during data analysis as well as the main limitations and advantages of this approach.
Keywords: Biosensor; Genetically encoded probe s; Lipid binding domain; Lipid signaling; Live imaging; Phosphatidic acid; Phosphatidylinositol phosphate; Phosphatidylserine; Phospholipase; PtdIns.
Figures
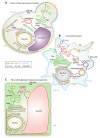


References
-
- Lemmon MA. Membrane recognition by phospholipid-binding domains. Nature reviews Molecular cell biology. 2008;9(2):99–111. doi:10.1038/nrm2328. - PubMed
-
- McLaughlin S, Murray D. Plasma membrane phosphoinositide organization by protein electrostatics. Nature. 2005;438(7068):605–611. doi:10.1038/nature04398. - PubMed
-
- Jean S, Kiger AA. Coordination between RAB GTPase and phosphoinositide regulation and functions. Nature reviews Molecular cell biology. 2012;13(7):463–470. doi:10.1038/nrm3379. - PubMed
-
- Kay JG, Grinstein S. Phosphatidylserine-mediated cellular signaling. Advances in experimental medicine and biology. 2013;991:177–193. doi:10.1007/978-94-007-6331-9_10. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

