Dysregulation of miR-212 Promotes Castration Resistance through hnRNPH1-Mediated Regulation of AR and AR-V7: Implications for Racial Disparity of Prostate Cancer
- PMID: 26553749
- PMCID: PMC7472565
- DOI: 10.1158/1078-0432.CCR-15-1606
Dysregulation of miR-212 Promotes Castration Resistance through hnRNPH1-Mediated Regulation of AR and AR-V7: Implications for Racial Disparity of Prostate Cancer
Abstract
Purpose: The causes of disproportionate incidence and mortality of prostate cancer among African Americans (AA) remain elusive. The purpose of this study was to investigate the mechanistic role and assess clinical utility of the splicing factor heterogeneous nuclear ribonucleoprotein H1 (hnRNP H1) in prostate cancer progression among AA men.
Experimental design: We employed an unbiased functional genomics approach coupled with suppressive subtractive hybridization (SSH) and custom cDNA microarrays to identify differentially expressed genes in microdissected tumors procured from age- and tumor grade-matched AA and Caucasian American (CA) men. Validation analysis was performed in independent cohorts and tissue microarrays. The underlying mechanisms of hnRNPH1 regulation and its impact on androgen receptor (AR) expression and tumor progression were explored.
Results: Aberrant coexpression of AR and hnRNPH1 and downregulation of miR-212 were detected in prostate tumors and correlate with disease progression in AA men compared with CA men. Ectopic expression of miR-212 mimics downregulated hnRNPH1 transcripts, which in turn reduced expression of AR and its splice variant AR-V7 (or AR3) in prostate cancer cells. hnRNPH1 physically interacts with AR and steroid receptor coactivator-3 (SRC-3) and primes activation of androgen-regulated genes in a ligand-dependent and independent manner. siRNA silencing of hnRNPH1 sensitized prostate cancer cells to bicalutamide and inhibited prostate tumorigenesis in vivo
Conclusions: Our findings define novel roles for hnRNPH1 as a putative oncogene, splicing factor, and an auxiliary AR coregulator. Targeted disruption of the hnRNPH1-AR axis may have therapeutic implications to improve clinical outcomes in patients with advanced prostate cancer, especially among AA men.
©2015 American Association for Cancer Research.
Conflict of interest statement
Disclosure of Potential Conflicts of Interest
No potential conflicts of interest were disclosed.
Figures


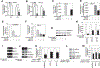



Similar articles
-
Galectin-3 Is Implicated in Tumor Progression and Resistance to Anti-androgen Drug Through Regulation of Androgen Receptor Signaling in Prostate Cancer.Anticancer Res. 2017 Jan;37(1):125-134. doi: 10.21873/anticanres.11297. Anticancer Res. 2017. PMID: 28011482
-
CX4945 suppresses the growth of castration-resistant prostate cancer cells by reducing AR-V7 expression.World J Urol. 2017 Aug;35(8):1213-1221. doi: 10.1007/s00345-016-1996-y. Epub 2017 Jan 19. World J Urol. 2017. PMID: 28105499
-
Insulin-Like Growth Factor 2 mRNA-Binding Protein 2 (IGF2BP2) Promotes Castration-Resistant Prostate Cancer Progression by Regulating AR-V7 mRNA Stability.Cancer Rep (Hoboken). 2025 Feb;8(2):e70096. doi: 10.1002/cnr2.70096. Cancer Rep (Hoboken). 2025. PMID: 39948708 Free PMC article.
-
Androgen receptors in hormone-dependent and castration-resistant prostate cancer.Pharmacol Ther. 2013 Dec;140(3):223-38. doi: 10.1016/j.pharmthera.2013.07.003. Epub 2013 Jul 13. Pharmacol Ther. 2013. PMID: 23859952 Review.
-
Androgen Receptor Splice Variant, AR-V7, as a Biomarker of Resistance to Androgen Axis-Targeted Therapies in Advanced Prostate Cancer.Clin Genitourin Cancer. 2020 Feb;18(1):1-10. doi: 10.1016/j.clgc.2019.09.015. Epub 2019 Sep 26. Clin Genitourin Cancer. 2020. PMID: 31653572 Review.
Cited by
-
Deregulated microRNAs Involved in Prostate Cancer Aggressiveness and Treatment Resistance Mechanisms.Cancers (Basel). 2023 Jun 10;15(12):3140. doi: 10.3390/cancers15123140. Cancers (Basel). 2023. PMID: 37370750 Free PMC article. Review.
-
The HNRNPF/H RNA binding proteins and disease.Wiley Interdiscip Rev RNA. 2023 Sep-Oct;14(5):e1788. doi: 10.1002/wrna.1788. Epub 2023 Apr 11. Wiley Interdiscip Rev RNA. 2023. PMID: 37042074 Free PMC article. Review.
-
Genetic and biological drivers of prostate cancer disparities in Black men.Nat Rev Urol. 2024 May;21(5):274-289. doi: 10.1038/s41585-023-00828-w. Epub 2023 Nov 14. Nat Rev Urol. 2024. PMID: 37964070 Review.
-
Advances of multi-omics applications in hepatic precancerous lesions and hepatocellular carcinoma: The role of extracellular vesicles.Front Mol Biosci. 2023 Mar 16;10:1114594. doi: 10.3389/fmolb.2023.1114594. eCollection 2023. Front Mol Biosci. 2023. PMID: 37006626 Free PMC article. Review.
-
MicroRNAs and Natural Compounds Mediated Regulation of TGF Signaling in Prostate Cancer.Front Pharmacol. 2021 Jan 27;11:613464. doi: 10.3389/fphar.2020.613464. eCollection 2020. Front Pharmacol. 2021. PMID: 33584291 Free PMC article. Review.
References
-
- Chen CD, Welsbie DS, Tran C, Baek SH, Chen R, Vessella R, et al. Molecular determinants of resistance to antiandrogen therapy. Nat Med 2014;10:33–9. - PubMed
-
- Heemers HV, Tindall DJ. Androgen receptor (AR) coregulators: a diversity of functions converging on and regulating the AR transcriptional complex. Endocr Rev 2007;28:778–808. - PubMed
-
- Siegel RL, Miller KD, Jemal A. Cancer Statistics. CA Cancer J Clin 2015;65: 5–29. - PubMed
-
- Powell IJ, Banerjee M, Novallo M, Sakr W, Grignon D, Wood DP, et al. Prostate cancer biochemical recurrence stage for stage is more frequent among African American than white men with locally advanced but not organ-confined disease. Urology 2000;55:246–51. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

