Unexpected Allosteric Network Contributes to LRH-1 Co-regulator Selectivity
- PMID: 26553876
- PMCID: PMC4714224
- DOI: 10.1074/jbc.M115.662874
Unexpected Allosteric Network Contributes to LRH-1 Co-regulator Selectivity
Abstract
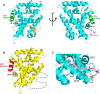
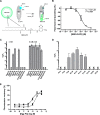
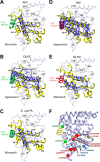
Phospholipids (PLs) are unusual signaling hormones sensed by the nuclear receptor liver receptor homolog-1 (LRH-1), which has evolved a novel allosteric pathway to support appropriate interaction with co-regulators depending on ligand status. LRH-1 plays an important role in controlling lipid and cholesterol homeostasis and is a potential target for the treatment of metabolic and neoplastic diseases. Although the prospect of modulating LRH-1 via small molecules is exciting, the molecular mechanism linking PL structure to transcriptional co-regulator preference is unknown. Previous studies showed that binding to an activating PL ligand, such as dilauroylphosphatidylcholine, favors LRH-1's interaction with transcriptional co-activators to up-regulate gene expression. Both crystallographic and solution-based structural studies showed that dilauroylphosphatidylcholine binding drives unanticipated structural fluctuations outside of the canonical activation surface in an alternate activation function (AF) region, encompassing the β-sheet-H6 region of the protein. However, the mechanism by which dynamics in the alternate AF influences co-regulator selectivity remains elusive. Here, we pair x-ray crystallography with molecular modeling to identify an unexpected allosteric network that traverses the protein ligand binding pocket and links these two elements to dictate selectivity. We show that communication between the alternate AF region and classical AF2 is correlated with the strength of the co-regulator interaction. This work offers the first glimpse into the conformational dynamics that drive this unusual PL-mediated nuclear hormone receptor activation.
Keywords: allosteric regulation; diabetes; lipids; molecular dynamics; nuclear receptor; phosphatidylcholine.
© 2016 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures






Similar articles
-
Antidiabetic phospholipid-nuclear receptor complex reveals the mechanism for phospholipid-driven gene regulation.Nat Struct Mol Biol. 2012 Apr 15;19(5):532-S2. doi: 10.1038/nsmb.2279. Nat Struct Mol Biol. 2012. PMID: 22504882 Free PMC article.
-
Divergent sequence tunes ligand sensitivity in phospholipid-regulated hormone receptors.J Biol Chem. 2013 Jul 12;288(28):20702-12. doi: 10.1074/jbc.M113.472837. Epub 2013 Jun 4. J Biol Chem. 2013. PMID: 23737522 Free PMC article.
-
Modulation of human nuclear receptor LRH-1 activity by phospholipids and SHP.Nat Struct Mol Biol. 2005 Apr;12(4):357-63. doi: 10.1038/nsmb910. Epub 2005 Feb 22. Nat Struct Mol Biol. 2005. PMID: 15723037
-
Allosteric controls of nuclear receptor function in the regulation of transcription.J Mol Biol. 2013 Jul 10;425(13):2317-29. doi: 10.1016/j.jmb.2013.03.017. Epub 2013 Mar 15. J Mol Biol. 2013. PMID: 23499886 Review.
-
Liver receptor homolog-1, an emerging metabolic modulator.Front Biosci. 2008 May 1;13:5950-8. doi: 10.2741/3128. Front Biosci. 2008. PMID: 18508634 Review.
Cited by
-
Development of a Versatile and Sensitive Direct Ligand Binding Assay for Human NR5A Nuclear Receptors.ACS Med Chem Lett. 2019 Nov 21;11(3):365-370. doi: 10.1021/acsmedchemlett.9b00442. eCollection 2020 Mar 12. ACS Med Chem Lett. 2019. PMID: 32184971 Free PMC article.
-
Solution Nuclear Magnetic Resonance Studies of the Ligand-Binding Domain of an Orphan Nuclear Receptor Reveal a Dynamic Helix in the Ligand-Binding Pocket.Biochemistry. 2018 Apr 3;57(13):1977-1986. doi: 10.1021/acs.biochem.8b00069. Epub 2018 Mar 22. Biochemistry. 2018. PMID: 29547262 Free PMC article.
-
International Union of Basic and Clinical Pharmacology CXIII: Nuclear Receptor Superfamily-Update 2023.Pharmacol Rev. 2023 Nov;75(6):1233-1318. doi: 10.1124/pharmrev.121.000436. Epub 2023 Aug 16. Pharmacol Rev. 2023. PMID: 37586884 Free PMC article. Review.
-
Insights into the mechanism of the PIK3CA E545K activating mutation using MD simulations.Sci Rep. 2018 Oct 19;8(1):15544. doi: 10.1038/s41598-018-27044-6. Sci Rep. 2018. PMID: 30341384 Free PMC article.
-
The nuclear receptor LRH-1 discriminates between ligands using distinct allosteric signaling circuits.Protein Sci. 2023 Oct;32(10):e4754. doi: 10.1002/pro.4754. Protein Sci. 2023. PMID: 37572334 Free PMC article.
References
-
- Walker A. K., Jacobs R. L., Watts J. L., Rottiers V., Jiang K., Finnegan D. M., Shioda T., Hansen M., Yang F., Niebergall L. J., Vance D. E., Tzoneva M., Hart A. C., and Näär A. M. (2011) A conserved SREBP-1/phosphatidylcholine feedback circuit regulates lipogenesis in metazoans. Cell 147, 840–852 - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

