Genome-Wide Estimates of Mutation Rates and Spectrum in Schizosaccharomyces pombe Indicate CpG Sites are Highly Mutagenic Despite the Absence of DNA Methylation
- PMID: 26564949
- PMCID: PMC4704713
- DOI: 10.1534/g3.115.022129
Genome-Wide Estimates of Mutation Rates and Spectrum in Schizosaccharomyces pombe Indicate CpG Sites are Highly Mutagenic Despite the Absence of DNA Methylation
Abstract
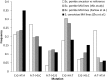
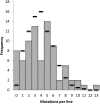
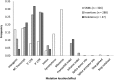
We accumulated mutations for 1952 generations in 79 initially identical, haploid lines of the fission yeast Schizosaccharomyces pombe, and then performed whole-genome sequencing to determine the mutation rates and spectrum. We captured 696 spontaneous mutations across the 79 mutation accumulation (MA) lines. We compared the mutation spectrum and rate to a recently published equivalent experiment on the same species, and to another model ascomycetous yeast, the budding yeast Saccharomyces cerevisiae. While the two species are approximately 600 million years diverged from each other, they share similar life histories, genome size and genomic G/C content. We found that Sc. pombe and S. cerevisiae have similar mutation rates, but Sc. pombe exhibits a stronger insertion bias. Intriguingly, we observed an increased mutation rate at cytosine nucleotides, specifically CpG nucleotides, which is also seen in S. cerevisiae. However, the absence of methylation in Sc. pombe and the pattern of mutation at these sites, primarily C → A as opposed to C → T, strongly suggest that the increased mutation rate is not caused by deamination of methylated cytosines. This result implies that the high mutability of CpG dinucleotides in other species may be caused in part by a methylation-independent mechanism. Many of our findings mirror those seen in the recent study, despite the use of different passaging conditions, indicating that MA is a reliable method for estimating mutation rates and spectra.
Keywords: cytosine mutation; insertion bias; mutation accumulation.
Copyright © 2016 Behringer and Hall.
Figures


References
-
- Andolfatto P., 2005. Adaptive evolution of non-coding DNA in Drosophila. Nature 437: 1149–1152. - PubMed
-
- Antequera F., Tamame M., Villanueva J., Santos T., 1984. DNA methylation in the fungi. J. Biol. Chem. 259: 8033–8036. - PubMed
-
- Aronesty, E., 2011 ea-utils: Command-line tools for processing biological sequencing data. Available at: http://code.google.com/p/ea-utils.
-
- Bell J., 2008. A simple way to treat PCR products prior to sequencing using ExoSAP-IT. Biotechniques 44: 834. - PubMed
-
- Bestor T. H., Verdine G. L., 1994. DNA methyltransferases. Curr. Opin. Cell Biol. 6: 380–389. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
