Non-Standard Genetic Codes Define New Concepts for Protein Engineering
- PMID: 26569314
- PMCID: PMC4695839
- DOI: 10.3390/life5041610
Non-Standard Genetic Codes Define New Concepts for Protein Engineering
Abstract
The essential feature of the genetic code is the strict one-to-one correspondence between codons and amino acids. The canonical code consists of three stop codons and 61 sense codons that encode 20% of the amino acid repertoire observed in nature. It was originally designated as immutable and universal due to its conservation in most organisms, but sequencing of genes from the human mitochondrial genomes revealed deviations in codon assignments. Since then, alternative codes have been reported in both nuclear and mitochondrial genomes and genetic code engineering has become an important research field. Here, we review the most recent concepts arising from the study of natural non-standard genetic codes with special emphasis on codon re-assignment strategies that are relevant to engineering genetic code in the laboratory. Recent tools for synthetic biology and current attempts to engineer new codes for incorporation of non-standard amino acids are also reviewed in this article.
Keywords: amino acids; biotechnology; codon reassignment; evolution; genetic code.
Figures


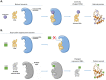
References
-
- Santos M., Santos M.A. Structural and molecular features of non-standard genetic codes. In: Cannarozzi G.M., Schneider A., editors. Codon Evolution: Mechanisms and Models. Oxford University Press; New York, NY, USA: 2012. pp. 258–271.
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

