A new network representation of the metabolism to detect chemical transformation modules
- PMID: 26573681
- PMCID: PMC4647279
- DOI: 10.1186/s12859-015-0809-4
A new network representation of the metabolism to detect chemical transformation modules
Abstract
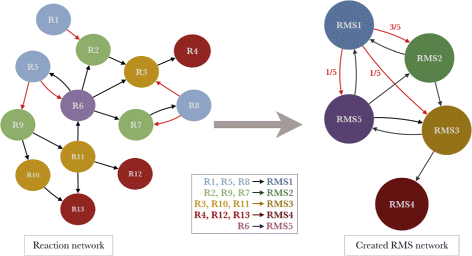
Background: Metabolism is generally modeled by directed networks where nodes represent reactions and/or metabolites. In order to explore metabolic pathway conservation and divergence among organisms, previous studies were based on graph alignment to find similar pathways. Few years ago, the concept of chemical transformation modules, also called reaction modules, was introduced and correspond to sequences of chemical transformations which are conserved in metabolism. We propose here a novel graph representation of the metabolic network where reactions sharing a same chemical transformation type are grouped in Reaction Molecular Signatures (RMS).
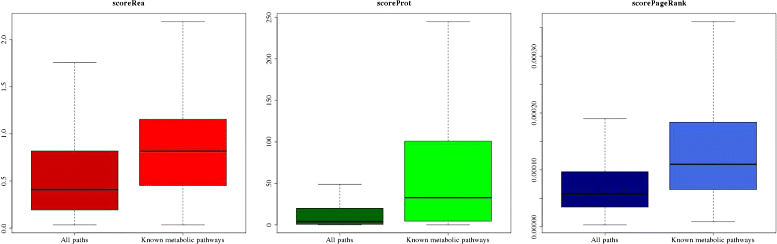
Results: RMS were automatically computed for all reactions and encode changes in atoms and bonds. A reaction network containing all available metabolic knowledge was then reduced by an aggregation of reaction nodes and edges to obtain a RMS network. Paths in this network were explored and a substantial number of conserved chemical transformation modules was detected. Furthermore, this graph-based formalism allows us to define several path scores reflecting different biological conservation meanings. These scores are significantly higher for paths corresponding to known metabolic pathways and were used conjointly to build association rules that should predict metabolic pathway types like biosynthesis or degradation.
Conclusions: This representation of metabolism in a RMS network offers new insights to capture relevant metabolic contexts. Furthermore, along with genomic context methods, it should improve the detection of gene clusters corresponding to new metabolic pathways.
Figures
Similar articles
-
Modular architecture of metabolic pathways revealed by conserved sequences of reactions.J Chem Inf Model. 2013 Mar 25;53(3):613-22. doi: 10.1021/ci3005379. Epub 2013 Feb 19. J Chem Inf Model. 2013. PMID: 23384306 Free PMC article.
-
Chemical and genomic evolution of enzyme-catalyzed reaction networks.FEBS Lett. 2013 Sep 2;587(17):2731-7. doi: 10.1016/j.febslet.2013.06.026. Epub 2013 Jun 28. FEBS Lett. 2013. PMID: 23816707 Review.
-
Development of a chemical structure comparison method for integrated analysis of chemical and genomic information in the metabolic pathways.J Am Chem Soc. 2003 Oct 1;125(39):11853-65. doi: 10.1021/ja036030u. J Am Chem Soc. 2003. PMID: 14505407
-
Inferring branching pathways in genome-scale metabolic networks.BMC Syst Biol. 2009 Oct 29;3:103. doi: 10.1186/1752-0509-3-103. BMC Syst Biol. 2009. PMID: 19874610 Free PMC article.
-
Networks and Graphs Discovery in Metabolomics Data Analysis and Interpretation.Front Mol Biosci. 2022 Mar 8;9:841373. doi: 10.3389/fmolb.2022.841373. eCollection 2022. Front Mol Biosci. 2022. PMID: 35350714 Free PMC article. Review.
Cited by
-
Metabolic pathway reconstruction strategies for central metabolism and natural product biosynthesis.Biophys Physicobiol. 2016 Jul 15;13:195-205. doi: 10.2142/biophysico.13.0_195. eCollection 2016. Biophys Physicobiol. 2016. PMID: 27924274 Free PMC article. Review.
References
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

