Ant colony optimisation of decision tree and contingency table models for the discovery of gene-gene interactions
- PMID: 26577156
- PMCID: PMC8687348
- DOI: 10.1049/iet-syb.2015.0017
Ant colony optimisation of decision tree and contingency table models for the discovery of gene-gene interactions
Abstract
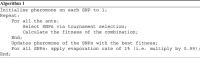
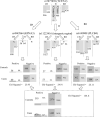
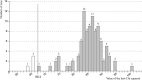
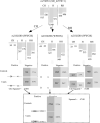
In this study, ant colony optimisation (ACO) algorithm is used to derive near-optimal interactions between a number of single nucleotide polymorphisms (SNPs). This approach is used to discover small numbers of SNPs that are combined into a decision tree or contingency table model. The ACO algorithm is shown to be very robust as it is proven to be able to find results that are discriminatory from a statistical perspective with logical interactions, decision tree and contingency table models for various numbers of SNPs considered in the interaction. A large number of the SNPs discovered here have been already identified in large genome-wide association studies to be related to type II diabetes in the literature, lending additional confidence to the results.
Figures


References
-
- Oki, N. : ‘On considering epistasis in genome wide association studies’ (North Carolina State University, North Carolina, USA: 2012)
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

