An experimentally supported model of the Bacillus subtilis global transcriptional regulatory network
- PMID: 26577401
- PMCID: PMC4670728
- DOI: 10.15252/msb.20156236
An experimentally supported model of the Bacillus subtilis global transcriptional regulatory network
Abstract
Organisms from all domains of life use gene regulation networks to control cell growth, identity, function, and responses to environmental challenges. Although accurate global regulatory models would provide critical evolutionary and functional insights, they remain incomplete, even for the best studied organisms. Efforts to build comprehensive networks are confounded by challenges including network scale, degree of connectivity, complexity of organism-environment interactions, and difficulty of estimating the activity of regulatory factors. Taking advantage of the large number of known regulatory interactions in Bacillus subtilis and two transcriptomics datasets (including one with 38 separate experiments collected specifically for this study), we use a new combination of network component analysis and model selection to simultaneously estimate transcription factor activities and learn a substantially expanded transcriptional regulatory network for this bacterium. In total, we predict 2,258 novel regulatory interactions and recall 74% of the previously known interactions. We obtained experimental support for 391 (out of 635 evaluated) novel regulatory edges (62% accuracy), thus significantly increasing our understanding of various cell processes, such as spore formation.
Keywords: Bacillus subtilis; network inference; sporulation; transcriptional networks.
© 2015 The Authors. Published under the terms of the CC BY 4.0 license.
Figures


Partial Pearson correlation between
mRNA transcription levels (PY 79 dataset) was computed for eachTF –target gene pair in theGS (top histogram). Partial Pearson correlation was also computed between the estimated activity of aTF and the transcription of its targets (bottom histogram).The advantage of estimating
TFA is illustrated for three regulators. Each point corresponds to the results of one microarray experiment, andTFA are estimated for each experimental condition. Top panel: A nonlinear correlation is observed between comK transcription and transcription of ComK targets, whereas a strong linear correlation is obtained between ComK activity and transcription of ComK targets. Middle panel: No correlation is observed between codY transcription and transcription of CodY targets. CodY activity is modulated byGTP and branched chain amino acids (BCAA ). A negative correlation is observed between estimated CodY activity and transcription of CodY targets. Bottom panel: Spo0A activity is modulated by phosphorylation. A better correlation is observed between Spo0A activity and transcription of Spo0A targets than between spo0A transcription and transcription of Spo0A targets.
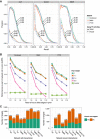
Precision–Recall plot of the confidence‐ranked interaction networks using
CLR , Genie3, and the Inferelator (no priors). Solid lines show performance usingTFA . Dashed lines show performance when noTFA are used (when raw expression values forTF s are used as predictors). The numbers superimposed on each curve indicate the area under the curve.Performance of
BBSR ‐TFA (AUPR : area under precision recall curve) on the combined,BSB 1 andPY 79, networks in the presence of false prior information. 50% of the edges in theGS are used as true priors, and various amounts of random edges are added. Performance is evaluated on the leave‐out set of interactions. Each point represents the median of five random samples of 50% of theGS set.Support from
KO data for the models predicted byBBSR ‐TFA , Genie3 (G3),CLR , and a consensus method (META ) that rank combines the prediction of the three methods. Methods were used without and withTFA (TFA tag). The number on top of each bar indicates the proportion of evaluated interactions withKO support for the corresponding method. Left and right panels show the support for each method when all interactions (recovered priors and novel interactions) and only novel interactions are considered, respectively.


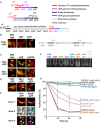
Genomic organization of the ytdA, yfnH, and spsI gene regions (ytdA is a paralog of yfnH and spsI). Putative gene functions are color‐coded. β score for each prediction is indicated in parenthesis.
Left: Fluorescence microscopy images for the YtdA‐
GFP fusion in sporulating cells in the indicated mutant backgrounds. Except where indicated otherwise, images were collected for sporulating cells at hour 6 after suspension in Sterlini–Mandelstam medium at 37°C. Right: Possible binding site for σK in the ytdA promoter. The consensus binding sequence for σK is also indicated (M is A or C).Spore coat localization of YtdA‐
GFP and SpsI‐GFP is dependent on cotE and independent of cotXYZ .Subcellular localization of YtcB‐
YFP , SpsJ‐YFP , and SpsK‐CFP during sporulation.Time course and dual labeling analysis of YfnH‐
YFP and SpsM‐CFP during sporulation.Spore adhesion to glass: A spsI deletion mutant and a spsI ytdA double mutant display strong adhesion.
Spore adhesion to hydrocarbons (hexadecane): A spsI deletion mutant and a spsI ytdA double mutant display strong adhesion. A spsI yfnH double mutant and a spsI yfnH ytdA triple mutant display intermediate adhesion. Error bars represent the standard deviation for three independent experiments.
References
Publication types
MeSH terms
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

