Rapid characterization of CRISPR-Cas9 protospacer adjacent motif sequence elements
- PMID: 26585795
- PMCID: PMC4653880
- DOI: 10.1186/s13059-015-0818-7
Rapid characterization of CRISPR-Cas9 protospacer adjacent motif sequence elements
Abstract
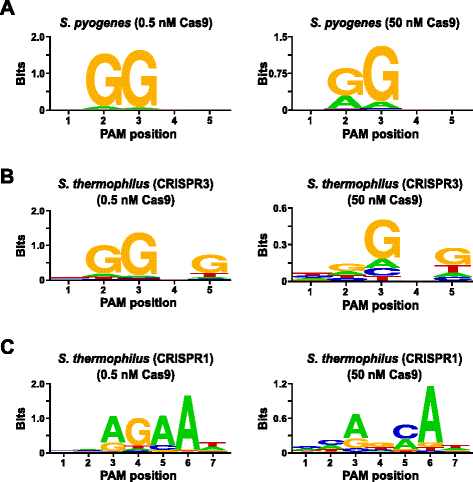
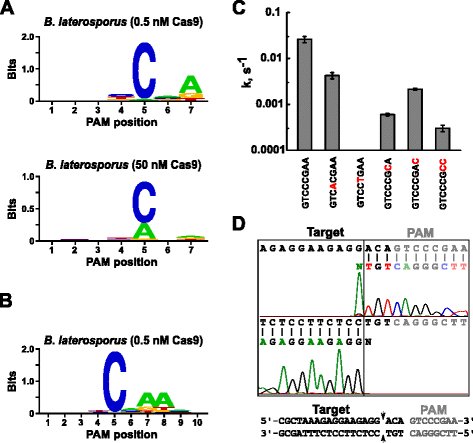
To expand the repertoire of Cas9s available for genome targeting, we present a new in vitro method for the simultaneous examination of guide RNA and protospacer adjacent motif (PAM) requirements. The method relies on the in vitro cleavage of plasmid libraries containing a randomized PAM as a function of Cas9-guide RNA complex concentration. Using this method, we accurately reproduce the canonical PAM preferences for Streptococcus pyogenes, Streptococcus thermophilus CRISPR3 (Sth3), and CRISPR1 (Sth1). Additionally, PAM and sgRNA solutions for a novel Cas9 protein from Brevibacillus laterosporus are provided by the assay and are demonstrated to support functional activity in vitro and in plants.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

