Activation of Type I and III Interferon Response by Mitochondrial and Peroxisomal MAVS and Inhibition by Hepatitis C Virus
- PMID: 26588843
- PMCID: PMC4654527
- DOI: 10.1371/journal.ppat.1005264
Activation of Type I and III Interferon Response by Mitochondrial and Peroxisomal MAVS and Inhibition by Hepatitis C Virus
Abstract
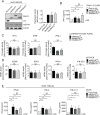
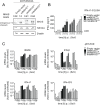
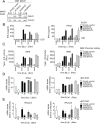
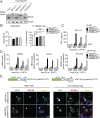
Sensing viruses by pattern recognition receptors (PRR) triggers the innate immune system of the host cell and activates immune signaling cascades such as the RIG-I/IRF3 pathway. Mitochondrial antiviral-signaling protein (MAVS, also known as IPS-1, Cardif, and VISA) is the crucial adaptor protein of this pathway localized on mitochondria, peroxisomes and mitochondria-associated membranes of the endoplasmic reticulum. Activation of MAVS leads to the production of type I and type III interferons (IFN) as well as IFN stimulated genes (ISGs). To refine the role of MAVS subcellular localization for the induction of type I and III IFN responses in hepatocytes and its counteraction by the hepatitis C virus (HCV), we generated various functional and genetic knock-out cell systems that were reconstituted to express mitochondrial (mito) or peroxisomal (pex) MAVS, exclusively. Upon infection with diverse RNA viruses we found that cells exclusively expressing pexMAVS mounted sustained expression of type I and III IFNs to levels comparable to cells exclusively expressing mitoMAVS. To determine whether viral counteraction of MAVS is affected by its subcellular localization we employed infection of cells with HCV, a major causative agent of chronic liver disease with a high propensity to establish persistence. This virus efficiently cleaves MAVS via a viral protease residing in its nonstructural protein 3 (NS3) and this strategy is thought to contribute to the high persistence of this virus. We found that both mito- and pexMAVS were efficiently cleaved by NS3 and this cleavage was required to suppress activation of the IFN response. Taken together, our findings indicate comparable activation of the IFN response by pex- and mitoMAVS in hepatocytes and efficient counteraction of both MAVS species by the HCV NS3 protease.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

