TRAIP promotes DNA damage response during genome replication and is mutated in primordial dwarfism
- PMID: 26595769
- PMCID: PMC4697364
- DOI: 10.1038/ng.3451
TRAIP promotes DNA damage response during genome replication and is mutated in primordial dwarfism
Abstract
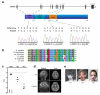
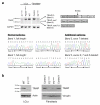
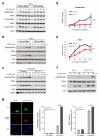
DNA lesions encountered by replicative polymerases threaten genome stability and cell cycle progression. Here we report the identification of mutations in TRAIP, encoding an E3 RING ubiquitin ligase, in patients with microcephalic primordial dwarfism. We establish that TRAIP relocalizes to sites of DNA damage, where it is required for optimal phosphorylation of H2AX and RPA2 during S-phase in response to ultraviolet (UV) irradiation, as well as fork progression through UV-induced DNA lesions. TRAIP is necessary for efficient cell cycle progression and mutations in TRAIP therefore limit cellular proliferation, providing a potential mechanism for microcephaly and dwarfism phenotypes. Human genetics thus identifies TRAIP as a component of the DNA damage response to replication-blocking DNA lesions.
Figures



References
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

