Large-Scale Computational Screening Identifies First in Class Multitarget Inhibitor of EGFR Kinase and BRD4
- PMID: 26596901
- PMCID: PMC4657038
- DOI: 10.1038/srep16924
Large-Scale Computational Screening Identifies First in Class Multitarget Inhibitor of EGFR Kinase and BRD4
Abstract
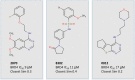
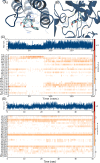
Inhibition of cancer-promoting kinases is an established therapeutic strategy for the treatment of many cancers, although resistance to kinase inhibitors is common. One way to overcome resistance is to target orthogonal cancer-promoting pathways. Bromo and Extra-Terminal (BET) domain proteins, which belong to the family of epigenetic readers, have recently emerged as promising therapeutic targets in multiple cancers. The development of multitarget drugs that inhibit kinase and BET proteins therefore may be a promising strategy to overcome tumor resistance and prolong therapeutic efficacy in the clinic. We developed a general computational screening approach to identify novel dual kinase/bromodomain inhibitors from millions of commercially available small molecules. Our method integrated machine learning using big datasets of kinase inhibitors and structure-based drug design. Here we describe the computational methodology, including validation and characterization of our models and their application and integration into a scalable virtual screening pipeline. We screened over 6 million commercially available compounds and selected 24 for testing in BRD4 and EGFR biochemical assays. We identified several novel BRD4 inhibitors, among them a first in class dual EGFR-BRD4 inhibitor. Our studies suggest that this computational screening approach may be broadly applicable for identifying dual kinase/BET inhibitors with potential for treating various cancers.
Figures






References
-
- Cohen P. Protein kinases—the major drug targets of the twenty-first century? Nat. Rev. Drug Discov. 1, 309–315 (2002). - PubMed
-
- Tsai C.-J. & Nussinov R. The molecular basis of targeting protein kinases in cancer therapeutics. In Semin. Cancer Biol. vol. 23, 235–242 (Academic Press, 2013). - PubMed
-
- Wu Q., Yang Z., Nie Y., Shi Y. & Fan D. Multi-drug resistance in cancer chemotherapeutics: Mechanisms and lab approaches. Cancer Lett. 347, 159–166 (2014). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

