Nanotechnologies for biomedical science and translational medicine
- PMID: 26598663
- PMCID: PMC4664315
- DOI: 10.1073/pnas.1515202112
Nanotechnologies for biomedical science and translational medicine
Abstract
In 2000 the United States launched the National Nanotechnology Initiative and, along with it, a well-defined set of goals for nanomedicine. This Perspective looks back at the progress made toward those goals, within the context of the changing landscape in biomedicine that has occurred over the past 15 years, and considers advances that are likely to occur during the next decade. In particular, nanotechnologies for health-related genomics and single-cell biology, inorganic and organic nanoparticles for biomedicine, and wearable nanotechnologies for wellness monitoring are briefly covered.
Keywords: biotechnology; nanomedicine; nanotechnology.
Conflict of interest statement
The author declares no conflict of interest.
Figures




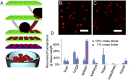


References
-
- National Science and Technology Council Committee on Technology. National Nanotechnology Initiative: leading to the next industrial revolution. A report by the Interagency Working Group on Nanoscale Science, Engineering, and Technology. Washington, DC: US Government; 2000. Available at: https://www.whitehouse.gov/files/documents/ostp/NSTC%20Reports/NNI2000.pdf.
-
- Gibbs WW. Medicine gets up close and personal. Nature. 2014;506(7487):144–145. - PubMed
-
- Leach DR, Krummel MF, Allison JP. Enhancement of antitumor immunity by CTLA-4 blockade. Science. 1996;271(5256):1734–1736. - PubMed
-
- Couzin-Frankel J. Breakthrough of the year 2013. Cancer immunotherapy. Science. 2013;342(6165):1432–1433. - PubMed
-
- Hayden EC. Technology: The $1,000 genome. Nature. 2014;507(7492):294–295. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

