Rheumatoid arthritis-associated RBPJ polymorphism alters memory CD4+ T cells
- PMID: 26604133
- PMCID: PMC4881829
- DOI: 10.1093/hmg/ddv474
Rheumatoid arthritis-associated RBPJ polymorphism alters memory CD4+ T cells
Abstract
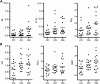
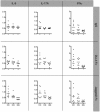
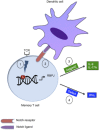
Notch signaling has recently emerged as an important regulator of immune responses in autoimmune diseases. The recombination signal-binding protein for immunoglobulin kappa J region (RBPJ) is a transcriptional repressor, but converts into a transcriptional activator upon activation of the canonical Notch pathway. Genome-wide association studies of rheumatoid arthritis (RA) identified a susceptibility locus, rs874040(CC), which implicated the RBPJ gene. Here, chromatin state mapping generated using the chromHMM algorithm reveals strong enhancer regions containing DNase I hypersensitive sites overlapping the rs874040 linkage disequilibrium block in human memory, but not in naïve CD4(+) T cells. The rs874040 overlapping this chromatin state was associated with increased RBPJ expression in stimulated memory CD4(+) T cells from healthy subjects homozygous for the risk allele (CC) compared with memory CD4(+) T cells bearing the protective allele (GG). Transcriptomic analysis of rs874040(CC) memory T cells showed a repression of canonical Notch target genes IL (interleukin)-9, IL-17 and interferon (IFN)γ in the basal state. Interestingly, activation of the Notch pathway using soluble Notch ligand, Jagged2-Fc, induced IL-9 and IL-17A while delta-like 4Fc, another Notch ligand, induced higher IFNγ expression in the rs874040(CC) memory CD4(+) T cells compared with their rs874040(GG) counterparts. In RA, RBPJ expression is elevated in memory T cells from RA patients compared with control subjects, and this was associated with induced inflammatory cytokines IL-9, IL-17A and IFNγ in response to Notch ligation in vitro. These findings demonstrate that the rs874040(CC) allele skews memory T cells toward a pro-inflammatory phenotype involving Notch signaling, thus increasing the susceptibility to develop RA.
© The Author 2015. Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oup.com.
Figures



Similar articles
-
Association of a Functional Single Nucleotide Polymorphism (rs874040) in the RBPJ Gene with Susceptibility to Rheumatoid Arthritis in Iranian Population.Avicenna J Med Biotechnol. 2021 Jul-Sep;13(3):166-170. doi: 10.18502/ajmb.v13i3.6368. Avicenna J Med Biotechnol. 2021. PMID: 34484646 Free PMC article.
-
Notch controls the survival of memory CD4+ T cells by regulating glucose uptake.Nat Med. 2015 Jan;21(1):55-61. doi: 10.1038/nm.3758. Epub 2014 Dec 15. Nat Med. 2015. PMID: 25501905
-
1,25-dihydroxyvitamin D3 modulates Th17 polarization and interleukin-22 expression by memory T cells from patients with early rheumatoid arthritis.Arthritis Rheum. 2010 Jan;62(1):132-42. doi: 10.1002/art.25043. Arthritis Rheum. 2010. PMID: 20039421
-
Transcription Factor RBPJ as a Molecular Switch in Regulating the Notch Response.Adv Exp Med Biol. 2021;1287:9-30. doi: 10.1007/978-3-030-55031-8_2. Adv Exp Med Biol. 2021. PMID: 33034023 Review.
-
Canonical and non-canonical Notch signaling in CD4⁺ T cells.Curr Top Microbiol Immunol. 2012;360:99-114. doi: 10.1007/82_2012_233. Curr Top Microbiol Immunol. 2012. PMID: 22695917 Review.
Cited by
-
Abnormal expression and mutation of the RBPJ gene may be involved in CD59- clonal proliferation in paroxysmal nocturnal hemoglobinuria.Exp Ther Med. 2019 Jun;17(6):4536-4546. doi: 10.3892/etm.2019.7475. Epub 2019 Apr 10. Exp Ther Med. 2019. PMID: 31086586 Free PMC article.
-
Association of a Functional Single Nucleotide Polymorphism (rs874040) in the RBPJ Gene with Susceptibility to Rheumatoid Arthritis in Iranian Population.Avicenna J Med Biotechnol. 2021 Jul-Sep;13(3):166-170. doi: 10.18502/ajmb.v13i3.6368. Avicenna J Med Biotechnol. 2021. PMID: 34484646 Free PMC article.
-
New insights into the phenomenon of remissions and relapses in autoimmune diseases and the puzzle of benign autoantibodies in healthy individuals.Front Immunol. 2025 May 9;16:1522356. doi: 10.3389/fimmu.2025.1522356. eCollection 2025. Front Immunol. 2025. PMID: 40416990 Free PMC article. Review.
-
RBPJ mediates uterine repair in the mouse and is reduced in women with recurrent pregnancy loss.FASEB J. 2018 May;32(5):2452-2466. doi: 10.1096/fj.201701032R. Epub 2018 Jan 8. FASEB J. 2018. PMID: 29242273 Free PMC article.
-
MS AHI1 genetic risk promotes IFNγ+ CD4+ T cells.Neurol Neuroimmunol Neuroinflamm. 2017 Dec 8;5(1):e414. doi: 10.1212/NXI.0000000000000414. eCollection 2018 Jan. Neurol Neuroimmunol Neuroinflamm. 2017. PMID: 29379820 Free PMC article.
References
-
- Begovich A.B., Carlton V.E., Honigberg L.A., Schrodi S.J., Chokkalingam A.P., Alexander H.C., Ardlie K.G., Huang Q., Smith A.M., Spoerke J.M. et al. (2004) A missense single-nucleotide polymorphism in a gene encoding a protein tyrosine phosphatase (PTPN22) is associated with rheumatoid arthritis. Am. J. Hum. Genet., 75, 330–337. - PMC - PubMed
-
- Plenge R.M., Padyukov L., Remmers E.F., Purcell S., Lee A.T., Karlson E.W., Wolfe F., Kastner D.L., Alfredsson L., Altshuler D. et al. (2005) Replication of putative candidate-gene associations with rheumatoid arthritis in >4000 samples from North America and Sweden: association of susceptibility with PTPN22, CTLA4, and PADI4. Am. J. Hum. Genet., 77, 1044–1060. - PMC - PubMed
-
- Gregersen P.K., Amos C.I., Lee A.T., Lu Y., Remmers E.F., Kastner D.L., Seldin M.F., Criswell L.A., Plenge R.M., Holers V.M. et al. (2009) REL, encoding a member of the NF-kappaB family of transcription factors, is a newly defined risk locus for rheumatoid arthritis. Nat. Genet., 41, 820–823. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

