The histone methyltransferase DOT1L: regulatory functions and a cancer therapy target
- PMID: 26609488
- PMCID: PMC4633909
The histone methyltransferase DOT1L: regulatory functions and a cancer therapy target
Abstract
DOT1L is a unique histone methyltransferase that targets the histone H3 lysine 79 (H3K79) residue for mono-, di- and tri- methylation. Histone H3K79 mono- and di-methylation results in active gene transcription, while H3K79 tri-methylation is associated with gene repression. DOT1L has a critical role in regulating gene transcription, development, cell cycle progression, somatic reprogramming and DNA damage repair. DOT1L interacts with Mixed Lineage Leukemia (MLL) fusion proteins, leading to enhanced H3K79 methylation, maintenance of open chromatin, overexpression of downstream oncogenes and leukemogenesis. Importantly, small molecule DOT1L inhibitors have been recently developed, and one of the DOT1L inhibitors is already under investigation in a Phase I clinical trial in patients with MLL fusion gene-driven leukemia.
Keywords: DOT1L; DOT1L inhibitors; gene transcription; histone methylation; mixed lineage leukemia.
Figures

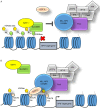
Similar articles
-
DOT1L/KMT4 recruitment and H3K79 methylation are ubiquitously coupled with gene transcription in mammalian cells.Mol Cell Biol. 2008 Apr;28(8):2825-39. doi: 10.1128/MCB.02076-07. Epub 2008 Feb 19. Mol Cell Biol. 2008. PMID: 18285465 Free PMC article.
-
The upstreams and downstreams of H3K79 methylation by DOT1L.Chromosoma. 2016 Sep;125(4):593-605. doi: 10.1007/s00412-015-0570-5. Epub 2016 Jan 4. Chromosoma. 2016. PMID: 26728620 Review.
-
DOT1L: a key target in normal chromatin remodelling and in mixed-lineage leukaemia treatment.Epigenetics. 2020 May;15(5):439-453. doi: 10.1080/15592294.2019.1699991. Epub 2019 Dec 28. Epigenetics. 2020. PMID: 31790636 Free PMC article. Review.
-
A medicinal chemistry perspective for targeting histone H3 lysine-79 methyltransferase DOT1L.J Med Chem. 2013 Nov 27;56(22):8972-83. doi: 10.1021/jm4007752. Epub 2013 Aug 14. J Med Chem. 2013. PMID: 23879463 Free PMC article. Review.
-
The emerging roles of DOT1L in leukemia and normal development.Leukemia. 2014 Nov;28(11):2131-8. doi: 10.1038/leu.2014.169. Epub 2014 May 23. Leukemia. 2014. PMID: 24854991 Review.
Cited by
-
Role of epigenetics in modulation of immune response at the junction of host-pathogen interaction and danger molecule signaling.Pathog Dis. 2016 Oct;74(7):ftw082. doi: 10.1093/femspd/ftw082. Epub 2016 Aug 18. Pathog Dis. 2016. PMID: 27542389 Free PMC article.
-
Insights Into the Properties, Biological Functions, and Regulation of USP21.Front Pharmacol. 2022 Jun 30;13:944089. doi: 10.3389/fphar.2022.944089. eCollection 2022. Front Pharmacol. 2022. PMID: 35846989 Free PMC article. Review.
-
Prognostic and therapeutic value of disruptor of telomeric silencing-1-like (DOT1L) expression in patients with ovarian cancer.J Hematol Oncol. 2017 Jan 23;10(1):29. doi: 10.1186/s13045-017-0400-8. J Hematol Oncol. 2017. PMID: 28114995 Free PMC article.
-
Histone and DNA Methylation as Epigenetic Regulators of DNA Damage Repair in Gastric Cancer and Emerging Therapeutic Opportunities.Cancers (Basel). 2023 Oct 13;15(20):4976. doi: 10.3390/cancers15204976. Cancers (Basel). 2023. PMID: 37894343 Free PMC article. Review.
-
Microarray-based Analysis of Genes, Transcription Factors, and Epigenetic Modifications in Lung Cancer Exposed to Nitric Oxide.Cancer Genomics Proteomics. 2020 Jul-Aug;17(4):401-415. doi: 10.21873/cgp.20199. Cancer Genomics Proteomics. 2020. PMID: 32576585 Free PMC article.
References
-
- Strahl B, Allis CD. The language of covalent histone modifications. Nature. 2000;403:41–45. - PubMed
-
- Leung K, Cheng VW, Mok SW, Tsui SK. The Involvement of DNA Methylation and Histone Modification on the Epigenetic Regulation of Embryonic Stem cells and Induced Pluripotent Stem Cells. Curr Stem Cell Res Ther. 2014;9:388–95. - PubMed
-
- Pérez-Cadahía B, Drobic B, Khan P, Shivashankar CC, Davie JR. Current understanding and importance of histone phosphorylation in regulating chromatin biology. Curr Opin Drug Discov Devel. 2010;13:613–622. - PubMed
LinkOut - more resources
Full Text Sources
