Metabolic control of methylation and acetylation
- PMID: 26629854
- PMCID: PMC4731252
- DOI: 10.1016/j.cbpa.2015.10.030
Metabolic control of methylation and acetylation
Abstract
Methylation and acetylation of DNA and histone proteins are the chemical basis for epigenetics. From bacteria to humans, methylation and acetylation are sensitive to cellular metabolic status. Modification rates depend on the availability of one-carbon and two-carbon substrates (S-adenosylmethionine, acetyl-CoA, and in bacteria also acetyl-phosphate). In addition, they are sensitive to demodification enzyme cofactors (α-ketoglutarate, NAD(+)) and structural analog metabolites that function as epigenetic enzyme inhibitors (e.g., S-adenosylhomocysteine, 2-hydroxyglutarate). Methylation and acetylation likely initially evolved to tailor protein activities in microbes to their metabolic milieu. While the extracellular environment of mammals is more tightly controlled, the combined impact of nutrient abundance and metabolic enzyme expression impacts epigenetics in mammals sufficiently to drive important biological outcomes such as stem cell fate and cancer.
Copyright © 2015 Elsevier Ltd. All rights reserved.
Figures
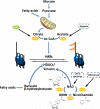

References
-
- Knight ZA, Shokat KM. Features of selective kinase inhibitors. Chem Biol. 2005;12:621–637. - PubMed
-
- Weinert BT, Iesmantavicius V, Wagner SA, Scholz C, Gummesson B, Beli P, Nystrom T, Choudhary C. Acetyl-phosphate is a critical determinant of lysine acetylation in E. coli. Mol Cell. 2013;51:265–272. [This paper shows that the main substrate driving lysine acetylation in E. coli is acetyl-phosphate, whose concentration impacts the overall extent of lysine acetylation. The NAD+-dependent deacetylase cobB removes the lysine acetylation.] - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

