Metagenomic Approach for Identification of the Pathogens Associated with Diarrhea in Stool Specimens
- PMID: 26637379
- PMCID: PMC4733167
- DOI: 10.1128/JCM.01965-15
Metagenomic Approach for Identification of the Pathogens Associated with Diarrhea in Stool Specimens
Abstract
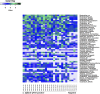
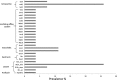
The potential to rapidly capture the entire microbial community structure and/or gene content makes metagenomic sequencing an attractive tool for pathogen identification and the detection of resistance/virulence genes in clinical settings. Here, we assessed the consistency between PCR from a diagnostic laboratory, quantitative PCR (qPCR) from a research laboratory, 16S rRNA gene sequencing, and metagenomic shotgun sequencing (MSS) for Clostridium difficile identification in diarrhea stool samples. Twenty-two C. difficile-positive diarrhea samples identified by PCR and qPCR and five C. difficile-negative diarrhea controls were studied. C. difficile was detected in 90.9% of C. difficile-positive samples using 16S rRNA gene sequencing, and C. difficile was detected in 86.3% of C. difficile-positive samples using MSS. CFU inferred from qPCR analysis were positively correlated with the relative abundance of C. difficile from 16S rRNA gene sequencing (r(2) = -0.60) and MSS (r(2) = -0.55). C. difficile was codetected with Clostridium perfringens, norovirus, sapovirus, parechovirus, and anellovirus in 3.7% to 27.3% of the samples. A high load of Candida spp. was found in a symptomatic control sample in which no causative agents for diarrhea were identified in routine clinical testing. Beta-lactamase and tetracycline resistance genes were the most prevalent (25.9%) antibiotic resistance genes in these samples. In summary, the proof-of-concept study demonstrated that next-generation sequencing (NGS) in pathogen detection is moderately correlated with laboratory testing and is advantageous in detecting pathogens without a priori knowledge.
Copyright © 2016, American Society for Microbiology. All Rights Reserved.
Figures

References
-
- Koser CU, Ellington MJ, Cartwright EJ, Gillespie SH, Brown NM, Farrington M, Holden MT, Dougan G, Bentley SD, Parkhill J, Peacock SJ. 2012. Routine use of microbial whole genome sequencing in diagnostic and public health microbiology. PLoS Pathog 8:e1002824. doi:10.1371/journal.ppat.1002824. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

