Radical-mediated ring contraction in the biosynthesis of 7-deazapurines
- PMID: 26643180
- PMCID: PMC4753066
- DOI: 10.1016/j.sbi.2015.11.005
Radical-mediated ring contraction in the biosynthesis of 7-deazapurines
Abstract
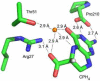
Pyrrolopyrimidine containing natural products are widely distributed in Nature. The biosynthesis of the 7-deazapurine moiety that is common to all pyrrolopyrimidines entails multiple steps, one of which is a complex radical-mediated ring contraction reaction catalyzed by CDG synthase. Herein we review the biosynthetic pathways of deazapurines, focusing on the biochemical and structural insights into CDG synthase.
Copyright © 2015 Elsevier Ltd. All rights reserved.
Figures

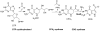

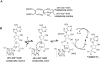


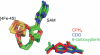
References
-
- Nishimura H, Katagiri K, Sato K, Mayama M, Shimoka N. Toyocamycin, a new anti-candida antibiotics. The Journal of antibiotics. 1956;9:60–62. - PubMed
-
- RajBhandary U, Chang HJ, Gross F, Harada F, Kimura S, Nishimura S. E. coli tyrosine transfer RNA-primary sequence and direct evidence for base pairing between terminal sequences. Fed Proc Fed Amer Soc Exp Biol. 1969;28:409.
-
- Harada F, Nishimura S. Possible anticodon sequences of tRNA His , tRNA Asn , and tRNA Asp from Escherichia coli B. Universal presence of nucleoside Q in the first postion of the anticondons of these transfer ribonucleic acids. Biochemistry. 1972;11:301–8. - PubMed
-
- Kasai H, Oashi Z, Harada F, Nishimura S, Oppenheimer NJ, Crain PF, Liehr JG, von Minden DL, McCloskey JA. Structure of the modified nucleoside Q isolated from Escherichia coli transfer ribonucleic acid. 7-(4,5-cis-Dihydroxy-1-cyclopenten-3-ylaminomethyl)-7-deazaguanosine. Biochemistry. 1975;14:4198–208. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

