Global Kinetic Analysis of Mammalian E3 Reveals pH-dependent NAD+/NADH Regulation, Physiological Kinetic Reversibility, and Catalytic Optimum
- PMID: 26644471
- PMCID: PMC4742739
- DOI: 10.1074/jbc.M115.676619
Global Kinetic Analysis of Mammalian E3 Reveals pH-dependent NAD+/NADH Regulation, Physiological Kinetic Reversibility, and Catalytic Optimum
Abstract
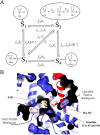
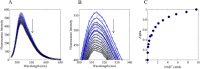
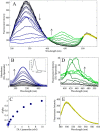
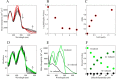
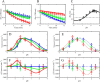
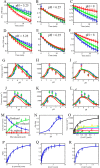
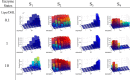
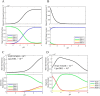
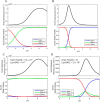
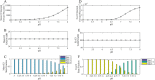
Mammalian E3 is an essential mitochondrial enzyme responsible for catalyzing the terminal reaction in the oxidative catabolism of several metabolites. E3 is a key regulator of metabolic fuel selection as a component of the pyruvate dehydrogenase complex (PDHc). E3 regulates PDHc activity by altering the affinity of pyruvate dehydrogenase kinase, an inhibitor of the enzyme complex, through changes in reduction and acetylation state of lipoamide moieties set by the NAD(+)/NADH ratio. Thus, an accurate kinetic model of E3 is needed to predict overall mammalian PDHc activity. Here, we have combined numerous literature data sets and new equilibrium spectroscopic experiments with a multitude of independently collected forward and reverse steady-state kinetic assays using pig heart E3. The latter kinetic assays demonstrate a pH-dependent transition of NAD(+) activation to inhibition, shown here, to our knowledge, for the first time in a single consistent data set. Experimental data were analyzed to yield a thermodynamically constrained four-redox-state model of E3 that simulates pH-dependent activation/inhibition and active site redox states for various conditions. The developed model was used to determine substrate/product conditions that give maximal E3 rates and show that, due to non-Michaelis-Menten behavior, the maximal flux is different compared with the classically defined kcat.
Keywords: enzyme kinetics; flavoprotein; global fitting; kcat; mathematical modeling; mitochondrial metabolism; pyruvate dehydrogenase complex (PDC).
© 2016 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures




References
-
- Perham R. N. (2000) Swinging arms and swinging domains in multifunctional enzymes: catalytic machines for multistep reactions. Annu. Rev. Biochem. 69, 961–1004 - PubMed
-
- Massey V. (1960) The identity of diaphorase and lipoyl dehydrogenase. Biochim. Biophys. Acta 37, 314–322 - PubMed
-
- Brautigam C. A., Chuang J. L., Tomchick D. R., Machius M., and Chuang D. T. (2005) Crystal structure of human dihydrolipoamide dehydrogenase: NAD+/NADH binding and the structural basis of disease-causing mutations. J. Mol. Biol. 350, 543–552 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

