Understanding Brassicaceae evolution through ancestral genome reconstruction
- PMID: 26653025
- PMCID: PMC4675067
- DOI: 10.1186/s13059-015-0814-y
Understanding Brassicaceae evolution through ancestral genome reconstruction
Erratum in
-
Erratum to: Understanding Brassicaceae evolution through ancestral genome reconstruction.Genome Biol. 2016 Apr 4;17:64. doi: 10.1186/s13059-016-0887-2. Genome Biol. 2016. PMID: 27044591 Free PMC article. No abstract available.
Abstract
Background: Brassicaceae is a family of green plants of high scientific and economic interest, including thale cress (Arabidopsis thaliana), cruciferous vegetables (cabbages) and rapeseed.
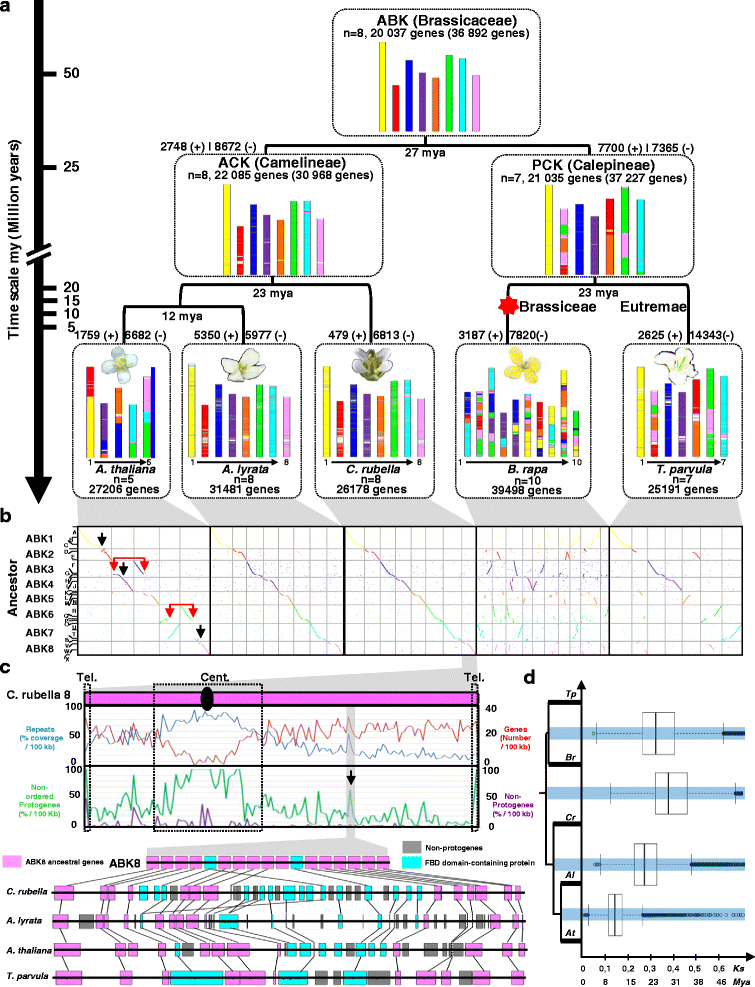
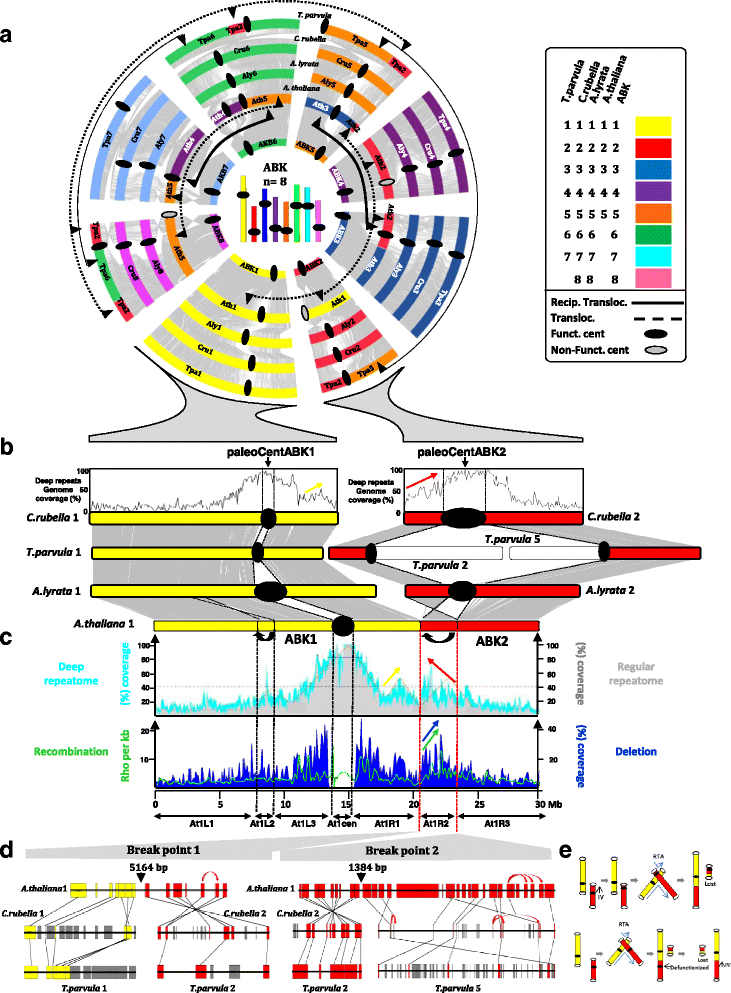
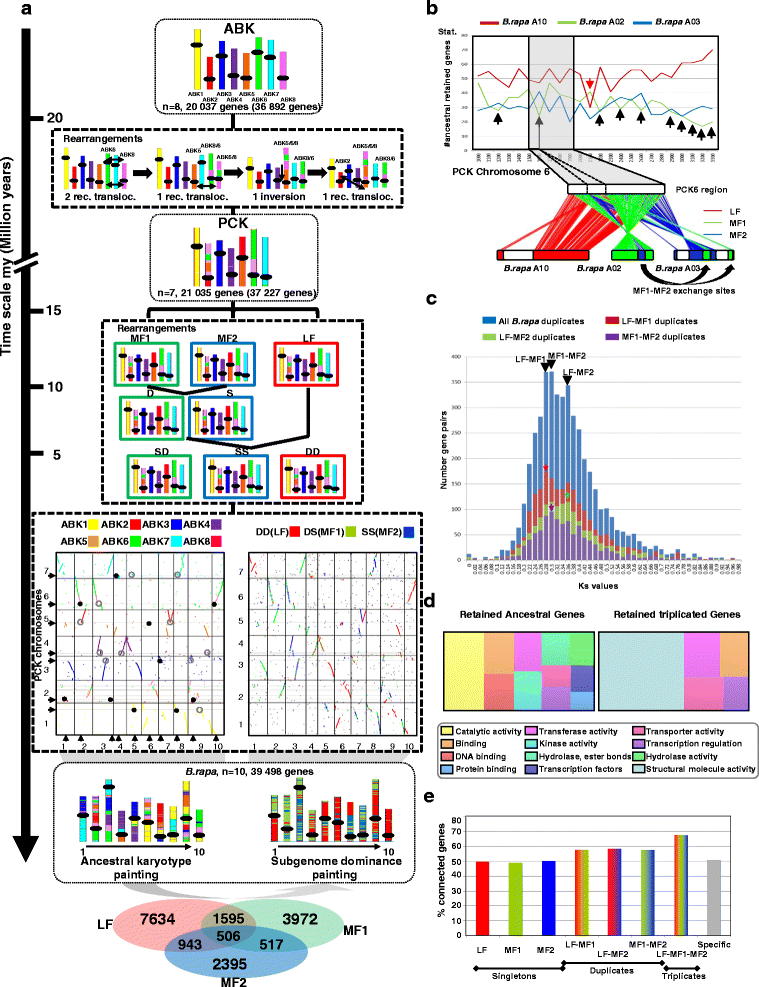
Results: We reconstruct an evolutionary framework of Brassicaceae composed of high-resolution ancestral karyotypes using the genomes of modern A. thaliana, Arabidopsis lyrata, Capsella rubella, Brassica rapa and Thellungiella parvula. The ancestral Brassicaceae karyotype (Brassicaceae lineages I and II) is composed of eight protochromosomes and 20,037 ordered and oriented protogenes. After speciation, it evolved into the ancestral Camelineae karyotype (eight protochromosomes and 22,085 ordered protogenes) and the proto-Calepineae karyotype (seven protochromosomes and 21,035 ordered protogenes) genomes.
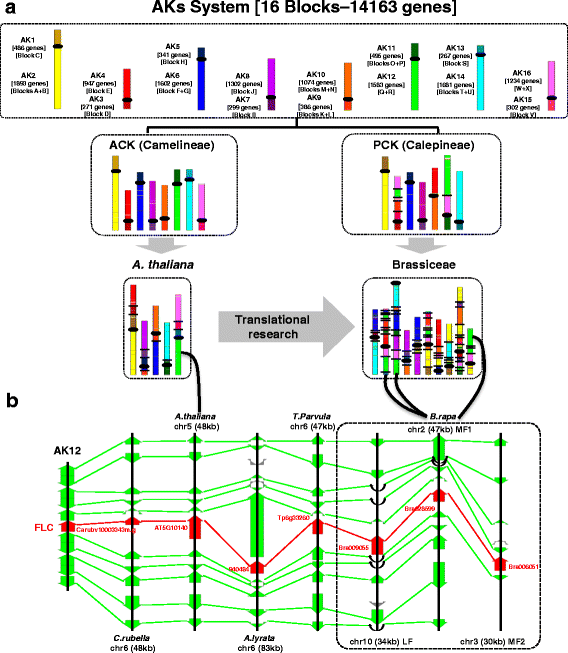
Conclusions: The three inferred ancestral karyotype genomes are shown here to be powerful tools to unravel the reticulated evolutionary history of extant Brassicaceae genomes regarding the fate of ancestral genes and genomic compartments, particularly centromeres and evolutionary breakpoints. This new resource should accelerate research in comparative genomics and translational research by facilitating the transfer of genomic information from model systems to species of agronomic interest.
Figures
Similar articles
-
The ABC's of comparative genomics in the Brassicaceae: building blocks of crucifer genomes.Trends Plant Sci. 2006 Nov;11(11):535-42. doi: 10.1016/j.tplants.2006.09.002. Epub 2006 Oct 6. Trends Plant Sci. 2006. PMID: 17029932 Review.
-
Karyotype and gene order evolution from reconstructed extinct ancestors highlight contrasts in genome plasticity of modern rosid crops.Genome Biol Evol. 2015 Jan 29;7(3):735-49. doi: 10.1093/gbe/evv014. Genome Biol Evol. 2015. PMID: 25637221 Free PMC article.
-
Deciphering the diploid ancestral genome of the Mesohexaploid Brassica rapa.Plant Cell. 2013 May;25(5):1541-54. doi: 10.1105/tpc.113.110486. Epub 2013 May 7. Plant Cell. 2013. PMID: 23653472 Free PMC article.
-
Chromosomal phylogeny and karyotype evolution in x=7 crucifer species (Brassicaceae).Plant Cell. 2008 Oct;20(10):2559-70. doi: 10.1105/tpc.108.062166. Epub 2008 Oct 3. Plant Cell. 2008. PMID: 18836039 Free PMC article.
-
Deciphering the evolutionary interplay between subgenomes following polyploidy: A paleogenomics approach in grasses.Am J Bot. 2016 Jul;103(7):1167-74. doi: 10.3732/ajb.1500459. Epub 2016 Jul 17. Am J Bot. 2016. PMID: 27425631 Review.
Cited by
-
Genome structural evolution in Brassica crops.Nat Plants. 2021 Jun;7(6):757-765. doi: 10.1038/s41477-021-00928-8. Epub 2021 May 27. Nat Plants. 2021. PMID: 34045706
-
Insights from the genomes of 4 diploid Camelina spp.G3 (Bethesda). 2022 Dec 1;12(12):jkac182. doi: 10.1093/g3journal/jkac182. G3 (Bethesda). 2022. PMID: 35976116 Free PMC article.
-
Reconstructing the genome of the most recent common ancestor of flowering plants.Nat Genet. 2017 Apr;49(4):490-496. doi: 10.1038/ng.3813. Epub 2017 Mar 13. Nat Genet. 2017. PMID: 28288112
-
Chromosome-level Thlaspi arvense genome provides new tools for translational research and for a newly domesticated cash cover crop of the cooler climates.Plant Biotechnol J. 2022 May;20(5):944-963. doi: 10.1111/pbi.13775. Epub 2022 Feb 6. Plant Biotechnol J. 2022. PMID: 34990041 Free PMC article.
-
Complex Interplay of Tandem, Segmental, Whole Genome Duplication, and Re-organization Drives Expansion of SAUR Gene Family in Brassicaceae.Biochem Genet. 2025 Jul 2. doi: 10.1007/s10528-025-11167-3. Online ahead of print. Biochem Genet. 2025. PMID: 40601104
References
-
- Warwick SI, Mummenhoff K, Sauder CA, Koch MA, Al-Shehbaz IA. Closing the gaps: phylogenetic relationships in the Brassicaceae based on DNA sequence data of nuclear ribosomal ITS region. Plant Systemat Evol. 2010;285:209–32. doi: 10.1007/s00606-010-0271-8. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

